|
|
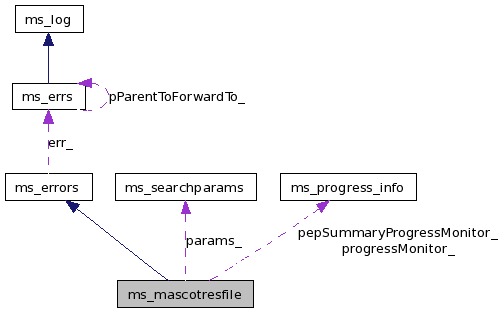
The first requirement before using any other functions or classes is to create an ms_mascotresfile object. More...
#include <ms_mascotresfile.hpp>

Public Types | |
| enum | err { ERR_NO_ERROR = 0x0000, ERR_NOMEM = 0x0001, ERR_NOSUCHFILE = 0x0002, ERR_READINGFILE = 0x0003, ERR_QUERYOUTOFRANGE = 0x0004, ERR_MISSINGENTRY = 0x0005, ERR_PEPSUMMPEPGET = 0x0006, ERR_PEPTIDESTR = 0x0007, ERR_ACCINPEPTIDESTR = 0x0008, ERR_PROTSUMM = 0x0009, ERR_PROTSUMMPEP = 0x000A, ERR_ADDPEPTIDES = 0x000B, ERR_MISSINGHIT = 0x000C, ERR_MISSINGSECTION = 0x000D, ERR_MISSINGSECTIONEND = 0x000E, ERR_MALFORMED_ERR_TOL = 0x000F, ERR_NO_ERR_TOL_PARENT = 0x0010, ERR_NULL_ACC_PEP_SUM = 0x0011, ERR_NULL_ACC_PROT_SUM = 0x0012, ERR_DUPE_ACCESSION = 0x0013, ERR_UNASSIGNED_PROG = 0x0014, ERR_UNASSIGNED_RANGE = 0x0015, ERR_UNASSIGNED_UNK = 0x0016, ERR_NO_UNIGENE_FILE = 0x0017, ERR_DUPLICATE_KEY = 0x0018, ERR_OLDRESULTSFILE = 0x0019, ERR_MALFORMED_TAG = 0x001A, ERR_MALFORMED_DRANGE = 0x001B, ERR_INVALID_NUMQUERIES = 0x001C, ERR_MALFORMED_TERMS = 0x001D, ERR_INVALID_RESFILE = 0x001E, ERR_INVALID_PROTDB = 0x001F, ERR_UNIGENE_MULTIDB = 0x0020, ERR_INVALID_CACHE_DIR = 0x0021, ERR_FAIL_OPEN_DAT_FILE = 0x0022, ERR_MISSING_CDB_FILE = 0x0023, ERR_FAIL_MK_CACHE_DIR = 0x0024, ERR_FAIL_MK_CDB_FILE = 0x0025, ERR_FAIL_CLOSE_FILE = 0x0026, ERR_FAIL_CDB_INIT = 0x0027, ERR_INVALID_CDB_FILE = 0x0028, ERR_WRITE_CDB_FILE = 0x0029, ERR_CDB_TOO_LARGE = 0x002A, ERR_NEED_64_BIT = 0x002B, ERR_CDB_64_BIT_REMAKE = 0x002C, ERR_CDB_OLD_VER_RETRY = 0x002D, ERR_CDB_OLD_VER_NO_RETRY = 0x002E, ERR_CDB_INCOMPLETE_RETRY = 0x002F, ERR_CDB_INCOMPLETE_NO_RETRY = 0x0030, ERR_CDB_BEING_CREATED = 0x0031, ERR_CDB_FAIL_REMOVE = 0x0032, ERR_CDB_FAIL_LOCK = 0x0033, ERR_CDB_FAIL_UNLOCK = 0x0034, ERR_CDB_SOURCE_CHANGE_RETRY = 0x0035, ERR_CDB_SOURCE_CHANGE_NO_RETRY = 0x0036, ERR_MISSING_PERCOLATOR_FILE = 0x0037, ERR_CANNOT_APPEND_RESFILE = 0x0038, ERR_CANNOT_APPEND_RESFILE_NO_FNAMES = 0x0039, ERR_RESULTS_NOT_CREATED = 0x003A } |
Definitions for error numbers. More... | |
| enum | FLAGS { RESFILE_NOFLAG = 0x00000000, RESFILE_USE_CACHE = 0x00000001, RESFILE_CACHE_IGNORE_ACC_DUPES = 0x00000002, RESFILE_USE_PARENT_PARAMS = 0x00000004, RESFILE_CACHE_IGNORE_DATE_CHANGE = 0x00000008 } |
Flags for opening the results file. More... | |
| enum | KA_TASK { KA_CREATEINDEX_CI = 0, KA_READFILE_RF = 1, KA_ASSIGNPROTEINS_AP = 2, KA_GROUPPROTEINS_GP = 3, KA_UNASSIGNEDLIST_UL = 4, KA_QUANTITATION = 5, KA_CREATECACHE_CC = 6, KA_THRESHFORFDR_FDR = 7, KA_LAST = 8 } |
Processing some results files is computationally intensive. These are the tasks that can be performed. More... | |
| enum | PERCOLATOR_FILE_NAMES { PERCOLATOR_INPUT_FILE = 0, PERCOLATOR_OUTPUT_TARGET = 1, PERCOLATOR_OUTPUT_DECOY = 2 } |
Offsets into a vector of Percolator filenames. More... | |
| enum | section { SEC_PARAMETERS, SEC_HEADER, SEC_MASSES, SEC_SUMMARY, SEC_MIXTURE, SEC_PEPTIDES, SEC_PROTEINS, SEC_QUERY1, SEC_QUANTITATION, SEC_UNIMOD, SEC_ENZYME, SEC_TAXONOMY, SEC_DECOYSUMMARY, SEC_DECOYMIXTURE, SEC_DECOYPEPTIDES, SEC_DECOYPROTEINS, SEC_ERRTOLSUMMARY, SEC_ERRTOLPEPTIDES, SEC_SPECTRAL_LIBRARY, SEC_LIBRARYPEPTIDES, SEC_LIBRARYSUMMARY, SEC_CROSSLINK_SUMMARY, SEC_CROSSLINK_PEPTIDES, SEC_CROSSLINKING, SEC_UNIMOD_XL, SEC_ERRTOLDECOYSUMMARY, SEC_ERRTOLDECOYPEPTIDES, SEC_INDEX, SEC_NUMSECTIONS } |
Section names in the standard mascot results files. More... | |
| enum | XML_SCHEMA { XML_SCHEMA_QUANTITATION = 0, XML_SCHEMA_UNIMOD = 1, XML_SCHEMA_DIRECTORY = 2, XML_SCHEMA_CROSSLINKING = 3, XML_SCHEMA_LAST = 4 } |
The results file contains sections in XML format and these need to be validated against a schema. More... | |
Public Member Functions | |
| ms_mascotresfile (const char *szFileName, const int keepAliveInterval=0, const char *keepAliveText="<!-- %d seconds -->\n", const unsigned int flags=RESFILE_NOFLAG, const char *cacheDirectory="../data/cache/%Y/%m", const char *XMLschemaDirectory=0, ms_progress_info *progressMonitor=0) | |
| Constructor to open a Mascot results file. | |
| bool | anyMSMS () const |
| Returns true if any of the queries in the search contain ions data. | |
| bool | anyPeptideSummaryMatches (const section sec=SEC_PEPTIDES) const |
| Returns true if there is a peptides section, and if there are any results in it. | |
| bool | anyPMF () const |
| Returns true if any of the queries in the search just contain a single peptide mass. | |
| bool | anySQ () const |
Returns true if any of the queries in the search contain seq or comp commands. | |
| bool | anyTag () const |
Returns true if any of the queries in the search contain tag or etag commands. | |
| void | appendErrors (ms_errs &errs) |
| Used internally. | |
| void | appendErrors (const ms_errors &src) |
| Copies all errors from another instance and appends them at the end of own list. | |
| int | appendResfile (const char *filename, int flags=RESFILE_USE_PARENT_PARAMS, const char *cacheDirectory=0) |
| Multiple results files can be summed together and treated as 'one'. | |
| void | clearAllErrors () |
| Remove all errors from the current list of errors. | |
| void | copyFrom (const ms_errors *right) |
| Use this member to make a copy of another instance. | |
| bool | doesSectionExist (const section sec) const |
| Returns true if there is an entry for the passed section. | |
| std::string | enumerateQuerySectionKeys (const int query, const int num, int *pPreviousNum=0, OFFSET64_T *pPreviousOffset=0) const |
| Get the key name for each item in a query section. | |
| std::string | enumerateSectionKeys (const section sec, const int num, int *pPreviousNum=0, OFFSET64_T *pPreviousOffset=0) const |
| Get the key name for each item in a section. | |
| std::string | get_ms_mascotresults_params (const ms_mascotoptions &opts, unsigned int *gpFlags, double *gpMinProbability, int *gpMaxHitsToReport, double *gpIgnoreIonsScoreBelow, unsigned int *gpMinPepLenInPepSummary, bool *gpUsePeptideSummary, unsigned int *gpFlags2) const |
| [Deprecated] Return default flags and parameters for creating an ms_peptidesummary or ms_proteinsummary object. | |
| std::string | get_ms_mascotresults_params (const ms_mascotoptions &opts, ms_mascotresults_params ¶ms) const |
| Return default flags and parameters for creating an ms_peptidesummary or ms_proteinsummary object. | |
| std::string | getCacheDirectory (bool processed=true) const |
| Returns the directory being used for cache files (if any). | |
| std::string | getCacheFileName () const |
| Returns the filename of the cache file. | |
| bool | getCrosslinkingMethod (ms_crosslinking_method *method) const |
| Return the crosslinking method object from the crosslinking section of the results file. | |
| DATABASE_TYPE | getDatabaseType (const int idx) const |
| Return database type if available. | |
| int | getDate () const |
| Returns the date and time of the search in seconds since midnight January 1st 1970. | |
| ms_mascotoptions::DECOY_ALGORITHM | getDecoyTypeForDB (const int idx=1) const |
| Returns the decoy algorithm type for a given database. | |
| bool | getEnzyme (ms_enzymefile *efile, const char *enzymeFileName=0) const |
Returns an object that represents enzyme-section as a reduced enzymes file. | |
| const ms_errs * | getErrorHandler () const |
| Retrive the error object using this function to get access to all errors and error parameters. | |
| int | getErrorNumber (const int num=-1) const |
| Return a specific error number - or ms_mascotresfile::ERR_NO_ERROR. | |
| std::string | getErrorString (const int num) const |
| Return a specific error as a string. | |
| int | getExecTime () const |
| Returns the time taken for the search. | |
| std::string | getFastaPath (int idx=1) const |
| Returns the path to the FASTA file used. | |
| std::string | getFastaVer (int idx=1) const |
| Returns the FASTA file version. | |
| std::string | getFileName (const int id=1) const |
| Returns the name of the results file passed into the constructor. | |
| int | getJobNumber (const int resfileID=1) const |
| Return the job number for this file - obtained from the file name. | |
| void | getKeepAlive (KA_TASK &kaTask, int &kaPercentage, std::string &kaAccession, int &kaHit, int &kaQuery, std::string &kaText) const |
| Return the progress indicators used by the keepAlive functions. | |
| int | getLastError () const |
| Return the last error number - or ms_mascotresfile::ERR_NO_ERROR. | |
| std::string | getLastErrorString () const |
| Return the last error number - or an empty string. | |
| std::string | getMascotVer () const |
| Returns the version of Mascot used to perform the search. | |
| bool | getMasses (ms_masses *masses) const |
| Returns an ms_masses object from the mass values in the results file. | |
| const ms_modification * | getMonoLinkModification (const int modNum, const int monoLink) const |
| Returns an ms_modification object that represents a monolink variable modification. | |
| std::string | getMSParserVersion () const |
| Returns the version number of the Mascot Parser library. | |
| int | getMultiFileQueryNumber (const int localQuery, const int fileId) const |
| Return the multi-file query number from the local query number in an appended file. | |
| int | getNumberOfErrors () const |
| Return the number of errors since the last call to clearAllErrors. | |
| int | getNumberOfResfiles () const |
| Multiple results files can be summed together and treated as 'one'. | |
| int | getNumEtSeqsSearched (const int idx=0) const |
| Returns the number of sequences searched in the second pass of an integrated error tolerant search. | |
| int | getNumHits (const section sec=SEC_SUMMARY) const |
| Returns the maximum number of hits possible for a protein summary. | |
| int | getNumLibraryEntries (const int idx=0) const |
| Returns the number of entries in the spectral library searched. | |
| int | getNumQueries (const int resfileID=0) const |
| Returns the number of queries (peptide masses or ms-ms spectra). | |
| double | getNumResidues (const int idx=0) const |
| Returns the number of residues in the FASTA file(s) searched. | |
| int | getNumSeqs (const int idx=0) const |
| Returns the number of sequences in the FASTA file(s) searched. | |
| int | getNumSeqsAfterTax (const int idx=0) const |
| Returns the number of sequences that passed the taxonomy filter in the FASTA file(s) searched. | |
| int | getObservedCharge (const int query, const bool decoy=false) const |
| The 'charge' returned will be 0 for Mr, otherwise it will be 1, -1, 2, -2, 3, -3 etc. and -100 for an error. | |
| double | getObservedIntensity (const int query) const |
| Returns the experimental intensity for the peptide. | |
| double | getObservedMass (const int query) const |
| Returns the experimental mass value as entered by the user. | |
| double | getObservedMrValue (const int query, const bool decoy=false) const |
| Returns the experimental mass value (as a relative mass) as entered by the user. | |
| std::vector< std::string > | getPercolatorFileNames () const |
| Retrieve the filenames use for percolator input and output. | |
| ms_progress_info * | getProgressInfo (bool forPeptideSummary=false) const |
| If a matrix_science::ms_progress_info object is passed to the constructor, this is returned here. | |
| bool | getQuantitation (ms_quant_configfile *qfile) const |
Returns an object that represents the quantitation section as a reduced quantitation.xml file. | |
| bool | getQuantitationMethod (ms_quant_method *qmethod) const |
| Return the quantitation method object from the quantitation section of the results file. | |
| int | getQuerySectionValue (const int query, const char *key, char *str, int maxLen) const |
| Return the string value from a query in the results file. | |
| double | getQuerySectionValueDouble (const int query, const char *key) const |
| Return the floating point value from a query in the results file. | |
| int | getQuerySectionValueInt (const int query, const char *key) const |
| Return the integer value from a query in the results file. | |
| std::string | getQuerySectionValueStr (const int query, const char *key) const |
| Return the string value from a query in the results file. | |
| int | getReferenceDatabaseNumberOfSL (const int idx) const |
| Return the database number of the reference database of a spectral library. | |
| std::string | getRepeatSearchString (const int query, const bool fullQuery=false) const |
| To perform a repeat search need to build up appropriate string. | |
| const ms_mascotresfile * | getResfile (int id) const |
| Returns a pointer to the resfile created by calling appendResfile. | |
| int | getSectionValue (const section sec, const char *key, char *str, int maxLen) const |
| Return the string value from any line in the results file. | |
| double | getSectionValueDouble (const section sec, const char *key) const |
| Return the floating point value from any line in the results file. | |
| int | getSectionValueInt (const section sec, const char *key) const |
| Return the integer value from any line in the results file. | |
| std::string | getSectionValueStr (const section sec, const char *key) const |
| Return the string value from any line in the results file. | |
| std::vector< int > | getSLDatabaseNumbersOfReference (const int idx) const |
| Return the database numbers of the spectral libraries whose reference database is at the given index. | |
| double | getSLFragmentTolerance (int idx=1) const |
| Returns the effective spectral library fragment tolerance. | |
| std::string | getSLFragmentToleranceUnit (int idx=1) const |
| Returns the unit of the effective spectral library fragment tolerance. | |
| bool | getSrcQueryAndFileIdForMultiFile (const int q, int &gsqNewQuery, int &gsqFileId) const |
| Return the query number and file ID in the source .dat file. | |
| bool | getTaxonomy (ms_taxonomyfile *tfile) const |
Returns an object that represents taxonomy-section as a reduced taxonomy file. | |
| bool | getUnimod (ms_umod_configfile *ufile, bool useSchemaFromResfile=false) const |
Returns an object that represents unimod-section as a reduced unimod_2.xml file. | |
| bool | getUnimodXL (ms_umod_configfile *ufile, bool useSchemaFromResfile=false) const |
Returns an object that represents unimod_xl-section as a reduced unimod_xl.xml file. | |
| std::string | getUniqueTaskID () const |
| Returns the unique task ID used by Mascot Daemon. | |
| std::string | getXMLschemaFilePath (XML_SCHEMA XMLschema) const |
| Gets the XML schema to be used by functions using quantitation or unimod. | |
| bool | isDatabaseTypeAvailable () const |
| Check whether database types are available. | |
| bool | isErrorTolerant () const |
| Returns true if the search was an error tolerant search. | |
| bool | isMSMS () const |
Returns true if the search was an MSMS search (SEARCH=MIS). | |
| bool | isPMF () const |
Returns true if the search was a PMF search (SEARCH=PMF). | |
| bool | isSQ () const |
Returns true if the search was a sequence query search (SEARCH=SQ). | |
| bool | isValid () const |
| Call this function to determine if there have been any errors. | |
| bool | outputKeepAlive () const |
| Outputs the "keep-alive" string during time-consuming operations. | |
| ms_searchparams & | params () const |
| Returns a reference to the search parameters class. | |
| void | resetKeepAlive (const int keepAliveInterval, const char *keepAliveText, const bool propagateToAppended=true, const bool resetStartTime=false) |
| Replace the existing keepAlive values with new values. | |
| void | setPercolatorFeatures (const char *percolatorFeatures, const char *additionalFeatures, const bool useRetentionTimes) |
| Set Percolator features before creating an ms_peptidesummary with Percolator scoring (deprecated). | |
| void | setPercolatorFeatures (const ms_mascotoptions &options, const char *additionalFeatures) |
| Set Percolator features before creating an ms_peptidesummary with Percolator scoring. | |
| bool | setXMLschemaFilePath (XML_SCHEMA XMLschema, const char *path) |
| Sets the XML schema to be used by functions using quantitation or unimod. | |
| bool | versionGreaterOrEqual (int major, int minor, int revision) const |
| Compare the value returned by getMascotVer() with the passed version number. | |
Static Public Member Functions | |
| static const char * | getSectionName (const section sec) |
| Return the section name as a string given the enum value. | |
| static bool | staticGetPercolatorFileNames (const char *szDatFileName, const char *cacheDirectory, const char *percolatorFeatures, const char *additionalFeatures, const bool useRetentionTimes, std::vector< std::string > &filenames, std::vector< bool > &exists) |
| Returns a list of the Percolator input and output files for the specified data file (deprecated). | |
| static bool | staticGetPercolatorFileNames (const char *szDatFileName, const char *cacheDirectory, const ms_mascotoptions &options, const char *additionalFeatures, std::vector< std::string > &filenames, std::vector< bool > &exists) |
| Returns a list of the Percolator input and output files for the specified data file. | |
| static bool | willCreateCache (const char *szFileName, const ms_mascotoptions &opts, const char *applicationName, std::string &resfileCacheFileName, unsigned int &cacheStatus) |
| Returns true if a cache file will be created when the ms_mascotresfile constructor is called. | |
| static bool | willCreateCache (const char *szFileName, const unsigned int flags, const char *cacheDirectory, std::string *cacheFileName) |
| Returns true if a cache file will be created when the ms_mascotresfile constructor is called. | |
The first requirement before using any other functions or classes is to create an ms_mascotresfile object.
You must create an object of this class before anything else. The constructor takes the file name as a parameter. Note that all key names are case insensitive.
peptide_list.cpp, repeat_search.cpp, resfile_error.cpp, resfile_info.cpp, resfile_input.cpp, resfile_params.cpp, and resfile_summary.cpp.
| enum err |
Definitions for error numbers.
See Using enumerated values and static const ints in Perl, Java, Python and C#. Messages are classified as fatal errors [F] or warnings [W]. A warning will not cause ms_mascotresfile::isValid() to return false.
| ERR_NO_ERROR |
[W] Success |
| ERR_NOMEM |
[F] Failed to allocate memory to load the file |
| ERR_NOSUCHFILE |
[F] The file passed in the constructor does not exist |
| ERR_READINGFILE |
[F] Opened the file successfully, but failed to read from it |
| ERR_QUERYOUTOFRANGE |
[F] Set if query < 1 or query > getNumQueries |
| ERR_MISSINGENTRY |
[F] Set if there is no qexp value in the file |
| ERR_PEPSUMMPEPGET |
[F] Value of q, p or h out of range, so cannot get peptide info |
| ERR_PEPTIDESTR |
[F] The string in the peptides block is not valid |
| ERR_ACCINPEPTIDESTR |
[F] Could not parse an item for a given accession in the peptide section |
| ERR_PROTSUMM |
[F] Error parsing a line in the protein summary |
| ERR_PROTSUMMPEP |
[F] Couldn't parse peptide information from the protein summary section |
| ERR_ADDPEPTIDES |
[F] Failed to add peptides when creating the peptide summary |
| ERR_MISSINGHIT |
[F] Missing hit in the summary section |
| ERR_MISSINGSECTION |
[F] Complete missing section in the file |
| ERR_MISSINGSECTIONEND |
[F] Missing end of section in the file |
| ERR_MALFORMED_ERR_TOL |
[W] Expecting a line of format: q1_p2_et_mods=0.984020,0.000000,Citrullination |
| ERR_NO_ERR_TOL_PARENT |
[F] No parent search file. See Error tolerant searches |
| ERR_NULL_ACC_PEP_SUM |
[W] An empty accession string has been found. Possible problem in database |
| ERR_NULL_ACC_PROT_SUM |
[W] An empty accession string has been found. Possible problem in database |
| ERR_DUPE_ACCESSION |
[W] A possible duplicate accession string has been found. Possible problem in database. |
| ERR_UNASSIGNED_PROG |
[F] Programming error! Calling getNumberOfUnassigned() or getUnassigned() before createUnassignedList() |
| ERR_UNASSIGNED_RANGE |
[F] Calling ms_mascotresults::getUnassigned() with out of range number |
| ERR_UNASSIGNED_UNK |
[F] Calling ms_mascotresults::getUnassigned() - unable to retrieve value |
| ERR_NO_UNIGENE_FILE |
[F] Failed to open the UniGene file specified |
| ERR_DUPLICATE_KEY |
[W] Duplicate entries with the same key in the named section. |
| ERR_OLDRESULTSFILE |
[F] Very old results file (last century!). Parser requires 1.02 or later |
| ERR_MALFORMED_TAG |
[W] Expecting a line in format: q1_p2_tag=1:3:5:6,2:4:12:6,... |
| ERR_MALFORMED_DRANGE |
[W] Expecting a line in format: q1_p2_drange=0,256 |
| ERR_INVALID_NUMQUERIES |
[W] Invalid number of queries in results file has been corrected. |
| ERR_MALFORMED_TERMS |
[W] Expecting a line in format: q1_p2_terms=A,B:-,I:... |
| ERR_INVALID_RESFILE |
[F] Invalid results file format - missing or corrupt headers |
| ERR_INVALID_PROTDB |
[W] Invalid h1_db-string format. Expecting an integer number. |
| ERR_UNIGENE_MULTIDB |
[W] UniGene index is not supported in multi-database search |
| ERR_INVALID_CACHE_DIR |
[F] Must specify a cache directory if using CDB cache files |
| ERR_FAIL_OPEN_DAT_FILE |
[F] Failed to open the results file for reading |
| ERR_MISSING_CDB_FILE |
[W] Cache file is missing or cannot be opened |
| ERR_FAIL_MK_CACHE_DIR |
[F] Failed to create cache directory for cache files |
| ERR_FAIL_MK_CDB_FILE |
[W] Failed to create an cache file |
| ERR_FAIL_CLOSE_FILE |
[W] Failed to close file |
| ERR_FAIL_CDB_INIT |
[W] Failed to initialise cache file (%s). Error code %d. |
| ERR_INVALID_CDB_FILE |
[W] Value in cdb cache file (%s) is corrupt: %s |
| ERR_WRITE_CDB_FILE |
[W] Failed to write to the cache file (%s). Error %d (%s) |
| ERR_CDB_TOO_LARGE |
[W] Cannot use cache file (s) which exceeded max size of s. Try deleting the cache file and retrying |
| ERR_NEED_64_BIT |
[F] This results file (%s) is too large for 32 bit Mascot Parser. Please upgrade to 64 bit. |
| ERR_CDB_64_BIT_REMAKE |
[W] Re-creating %s. Was too large for 32 bit, but may succeed with 64 bit |
| ERR_CDB_OLD_VER_RETRY |
[W] Cache file %s is an old version. Creating new cache file |
| ERR_CDB_OLD_VER_NO_RETRY |
[W] Cache file %s is an old version. Continuing without cache |
| ERR_CDB_INCOMPLETE_RETRY |
[W] Cache file %s was not complete. Re-creating the cache file |
| ERR_CDB_INCOMPLETE_NO_RETRY |
[W] Cache file %s was not complete. Continuing without cache |
| ERR_CDB_BEING_CREATED |
[W] Cache file %s being created by another task. Continuing without cache |
| ERR_CDB_FAIL_REMOVE |
[W] Failed to remove old cache file %s - error %s. Continuing without cache |
| ERR_CDB_FAIL_LOCK |
[W] Failed to lock cache file %s. Error code: %d |
| ERR_CDB_FAIL_UNLOCK |
[W] Failed to unlock cache file %s. Error code: %d |
| ERR_CDB_SOURCE_CHANGE_RETRY |
[W] %s changed. %s (was %s), %s bytes (was %s). Re-creating the cache file |
| ERR_CDB_SOURCE_CHANGE_NO_RETRY |
[W] %s changed. %s (was %s), %s bytes (was %s). Continuing without cache |
| ERR_MISSING_PERCOLATOR_FILE |
[F] Percolator file %s is missing. Cannot continue |
| ERR_CANNOT_APPEND_RESFILE |
[F] The file %s cannot be appended to %s because %s values are different |
| ERR_CANNOT_APPEND_RESFILE_NO_FNAMES |
[F] The file cannot be appended because %s values are different |
| ERR_RESULTS_NOT_CREATED |
[W] Attempting to call function %s before createSummary() has completed. |
| enum FLAGS |
Flags for opening the results file.
See Using enumerated values and static const ints in Perl, Java, Python and C# and Caching Mascot Results.
| RESFILE_NOFLAG |
The standard original functionality. Read the whole file into memory. |
| RESFILE_USE_CACHE |
Create the cache if it doesn't already exist. Use the cache rather than reading the whole .dat file into memory. |
| RESFILE_CACHE_IGNORE_ACC_DUPES |
When creating a cache file, don't check for duplicate accessions in the SEC_PROTEINS and SEC_DECOYPROTEINS sections which can save some time. Strongly recommend that this flag is never used unless performance becomes a real issue and it is known that ms_mascotoptions::getIgnoreDupeAccession was not defined for the relevant database(s) when they were compressed. |
| RESFILE_USE_PARENT_PARAMS |
For use when Combining multiple .dat files. The flags and parameters are then inherited from the parent search. |
| RESFILE_CACHE_IGNORE_DATE_CHANGE |
Opening the resfile cache CDB file should ignore the last modified timestamp on the .dat file. |
| enum KA_TASK |
Processing some results files is computationally intensive. These are the tasks that can be performed.
See Using enumerated values and static const ints in Perl, Java, Python and C#.
Used with getKeepAlive(), but also see outputKeepAlive()
| KA_CREATEINDEX_CI |
Creating a cache file when Using the ms_mascotresfile cache files. |
| KA_READFILE_RF |
Reading the results file into memory when not using a cache. |
| KA_ASSIGNPROTEINS_AP |
Assigning peptides to proteins to get a list of all possible proteins. |
| KA_GROUPPROTEINS_GP |
Grouping proteins using ms_mascotresults::MSRES_GROUP_PROTEINS or ms_mascotresults::MSRES_CLUSTER_PROTEINS. |
| KA_UNASSIGNEDLIST_UL |
Creating the unassigned list - see ms_mascotresults::createUnassignedList. |
| KA_QUANTITATION |
Calculating quantitation values for reporter and multiplex protocols. |
| KA_CREATECACHE_CC |
Creating a cache file when Using the ms_peptidesummary cache. |
| KA_THRESHFORFDR_FDR |
Calls to ms_mascotresults::getThresholdForFDRAboveHomology can be slow. |
| KA_LAST |
Placeholder that is equal to the number of possible tasks. |
Offsets into a vector of Percolator filenames.
See Using enumerated values and static const ints in Perl, Java, Python and C#.
Used with getPercolatorFileNames().
| enum section |
Section names in the standard mascot results files.
See Using enumerated values and static const ints in Perl, Java, Python and C#.
| SEC_PARAMETERS |
parameters section |
| SEC_HEADER |
header section |
| SEC_MASSES |
masses section |
| SEC_SUMMARY |
summary section |
| SEC_MIXTURE |
mixture section (pmf mixture) |
| SEC_PEPTIDES |
peptides section |
| SEC_PROTEINS |
proteins section |
| SEC_QUERY1 |
query1 section. Don't use, see getQuerySectionValueStr() etc. |
| SEC_QUANTITATION |
quantitation section |
| SEC_UNIMOD |
unimod section |
| SEC_ENZYME |
enzyme section |
| SEC_TAXONOMY |
taxonomy section |
| SEC_DECOYSUMMARY |
decoy_summary section. See also Target-decoy searches and false discovery rate. |
| SEC_DECOYMIXTURE |
decoy_mixture section. See also Target-decoy searches and false discovery rate. |
| SEC_DECOYPEPTIDES |
decoy_peptides section. See also Target-decoy searches and false discovery rate. |
| SEC_DECOYPROTEINS |
decoy_proteins section. See also Target-decoy searches and false discovery rate. |
| SEC_ERRTOLSUMMARY |
error tolerant summary section. See also Error tolerant searches. |
| SEC_ERRTOLPEPTIDES |
error tolerant peptides section. See also Error tolerant searches. |
| SEC_SPECTRAL_LIBRARY |
spectral library section. See also Spectral libraries. |
| SEC_LIBRARYPEPTIDES |
spectral library peptides section. See also Spectral libraries. |
| SEC_LIBRARYSUMMARY |
spectral library summary section. See also Spectral libraries. |
| SEC_CROSSLINK_SUMMARY |
crosslink_summary section. See also Crosslinked search results. |
| SEC_CROSSLINK_PEPTIDES |
crosslink_peptides section. See also Crosslinked search results. |
| SEC_CROSSLINKING |
crosslinking section |
| SEC_UNIMOD_XL |
unimod_xl section |
| SEC_ERRTOLDECOYSUMMARY |
error tolerant decoy summary section. See also Error tolerant searches. |
| SEC_ERRTOLDECOYPEPTIDES |
error tolerant decoy peptides section. See also Error tolerant searches. |
| SEC_INDEX |
index section |
| SEC_NUMSECTIONS |
!!don't use - place holder |
| enum XML_SCHEMA |
The results file contains sections in XML format and these need to be validated against a schema.
This is the list of schema that can be set using setXMLschemaFilePath() and getXMLschemaFilePath()
| XML_SCHEMA_QUANTITATION |
From the "quantation" mime section of the file. Valid aliases are: "http://www.matrixscience.com/xmlns/schema/quantitation_1" and "http://www.matrixscience.com/xmlns/schema/quantitation_2". |
| XML_SCHEMA_UNIMOD |
From the "unimod" mime section of the file. Valid alias is: http://www.unimod.org/xmlns/schema/unimod_2. |
| XML_SCHEMA_DIRECTORY |
From the value of XMLschemaDirectory passed into the ms_mascotresfile constructor. |
| XML_SCHEMA_CROSSLINKING |
From the "crosslinking" mime section of the file. Valid alias is: http://www.matrixscience.com/xmlns/schema/crosslinking_1. |
| XML_SCHEMA_LAST |
Placeholder that is equal to the number of possible schema. |
| ms_mascotresfile | ( | const char * | szFileName, |
| const int | keepAliveInterval = 0, |
||
| const char * | keepAliveText = "<!-- %d seconds -->\n", |
||
| const unsigned int | flags = RESFILE_NOFLAG, |
||
| const char * | cacheDirectory = "../data/cache/%Y/%m", |
||
| const char * | XMLschemaDirectory = 0, |
||
| ms_progress_info * | progressMonitor = 0 |
||
| ) |
Constructor to open a Mascot results file.
If the ms_mascotresfile::RESFILE_NOFLAG flag is specified, then the constructor reads the whole file into memory, and makes an in memory index of all the keys for fast lookup later. This was the default (and only) behaviour for Mascot Parser versions 2.2 and earlier.
If the ms_mascotresfile::RESFILE_USE_CACHE flag is specified, then the file is not read into memory, but a separate cache file containing offsets is used to read each line in the results file when requested. This is considerably faster if just a few lines of the results file need to be accessed and also takes less memory. The filenames for the cache file will be created by extracting the filename part from the path supplied in szFileName and appending '.cdb'. So, typically the cache filenames will be of the form Fxxxxx.dat.cdb. See Caching Mascot Results.
For HTML reports with large result files it is sometimes necessary to output HTML fragments to keep the connection alive. This can be done by specifying the interval at which the text is output (keepAliveInterval) and the text that should be output (keepAliveText). See outputKeepAlive() for more details.
The functions isValid() or getLastError() should be called after creating the object to determine if the file was valid and loaded properly.
Possible error values:
| szFileName | is the path to a valid Mascot results file |
| keepAliveInterval | is the interval in seconds between each time the keepAliveText is output to stdout. If a value of zero is specified, then no keep alive text will be output. |
| keepAliveText | is output every keepAliveInterval seconds while the file is being loaded. See outputKeepAlive() for further details. |
| flags | are created by bitwise ORing the ms_mascotresfile::FLAGS |
| cacheDirectory | is the location where any cache files are stored. See Specifying cache file directory. If cacheDirectory is null or empty and RESFILE_USE_CACHE is specified, then ERR_INVALID_CACHE_DIR will be set and the object will be invalid. Most applications should get this value from ms_mascotoptions::getCacheDirectory(). |
| XMLschemaDirectory | is the location where the xml schema files are located. Some sections of the results file are encoded in XML, and Mascot Parser needs to verify that these are in the correct format using 'xsd' schema files. The required files are supplied with Mascot Parser in the config directory. Either supply the path to that directory, or copy the .xsd files from it to another directory and supply the path to that (only the xsd files from that directory are required). If this parameter is 0 (the default), then it is assumed that the software is running on the Mascot Server and that the files are located in separate directories under ../html/xmlns/schema as defined in setXMLschemaFilePath(). The constructor does not check that the schema files exist, but subsequent calls to functions such as getUnimod() and getQuantitationMethod() will fail if the schema is not available. The function setXMLschemaFilePath() may be called to override this value passed in the constructor for individual schema files. |
| progressMonitor | is an optional parameter that can be used to track progress of the creation of this object. It can also be used (from another thread) to cancel the creation of the object by calling ms_progress_info::setBreak. If the passed progressMonitor has a subTask, created by calling ms_progress_info::addSubtask("Creating peptide summary", 100), then that subtask will be used when creating the ms_peptidesummary object. If the subtask doesn't exist, then one is created. Both pointers can be obtained by calling ms_mascotresfile::getProgressInfo(). Make sure that the calling function does not delete the object pointed to by this paramater until after the ms_mascotresfile object has been deleted. See: Maintaining object references: two rules of thumb |
| bool anyMSMS | ( | ) | const |
Returns true if any of the queries in the search contain ions data.
See also the isMSMS() member, although this function is the preferred one.
| bool anyPeptideSummaryMatches | ( | const section | sec = SEC_PEPTIDES ) |
const |
Returns true if there is a peptides section, and if there are any results in it.
If no parameter is passed to this function, information is returned about the standard peptides section.
This function is useful if, for example, you want to know if there is a peptides, decoy_peptides or et_peptides section in the results file. Very early versions of Mascot didn't have a peptides section for MS-MS searches, and the decoy and error tolerant peptides sections were added in Mascot 2.2, so it is safest to check that this function returns true before creating an ms_peptidesummary report. If this function returns false, consider creating an ms_proteinsummary instead.
This function will also return false if there is a peptides section that doesn't contain any results.
Even if this function returns true, it is possible that an ms_peptidesummary object created from the file will be empty if, for example, all the matches have ions scores below the specified minimum ions score.
| bool anyPMF | ( | ) | const |
Returns true if any of the queries in the search just contain a single peptide mass.
See also the isPMF() member, although this function is the preferred one.
| bool anySQ | ( | ) | const |
Returns true if any of the queries in the search contain seq or comp commands.
See also the isSQ() member, although this function is the preferred one.
seq or comp commands | bool anyTag | ( | ) | const |
| void appendErrors | ( | const ms_errors & | src ) | [inherited] |
Copies all errors from another instance and appends them at the end of own list.
| src | The object to copy the errors across from. See Maintaining object references: two rules of thumb. |
| int appendResfile | ( | const char * | filename, |
| int | flags = RESFILE_USE_PARENT_PARAMS, |
||
| const char * | cacheDirectory = 0 |
||
| ) |
Multiple results files can be summed together and treated as 'one'.
See Combining multiple .dat files.
Attempts to load the specifed results file and append to the existing file.
Any warning or error messages in the file are also appended to the existing object. If isValid() for the new file returns false, it is not appended and this function returns 0.
ms_mascotresfile::ERR_CANNOT_APPEND_RESFILE_NO_FNAMES and ms_mascotresfile::ERR_CANNOT_APPEND_RESFILE will be set if the file cannot be appended because of different parameters, such as a different enzyme.
| filename | is the path to the results file to append. |
| flags | are one of the ms_mascotresfile::FLAGS. If RESFILE_USE_PARENT_PARAMS is specified, then the flags, keepAlive and cache directory are copied from the parent object. |
| cacheDirectory | is the directory for the cache files if RESFILE_USE_CACHE has been specified. If RESFILE_USE_PARENT_PARAMS is specified and cacheDirectory is null or an empty string, then the cache directory for the parent object is used. |
| void clearAllErrors | ( | ) | [inherited] |
Remove all errors from the current list of errors.
The list of 'errors' can include fatal errors, warning messages, information messages and different levels of debugging messages.
All messages are accumulated into a list in this object, until clearAllErrors() is called.
See Error Handling.
| void copyFrom | ( | const ms_errors * | right ) | [inherited] |
Use this member to make a copy of another instance.
| right | is the source to initialise from |
| bool doesSectionExist | ( | const section | sec ) | const |
Returns true if there is an entry for the passed section.
This function is useful if, for example, you want to know if there is a peptides section in the results file. Very early versions of Mascot didn't have a peptides section for MS-MS searches.
| sec | is the section number |
| std::string enumerateQuerySectionKeys | ( | const int | query, |
| const int | num, | ||
| int * | pPreviousNum = 0, |
||
| OFFSET64_T * | pPreviousOffset = 0 |
||
| ) | const |
Get the key name for each item in a query section.
Enumerate through all the entries in a section. It returns the key name (not value) for a single item.
In version of Mascot Parser prior to version 2.3, the items are returned in alphabetical order. In Mascot Parser version 2.3 and later, the items are returned in the order in the results file.
See ms_searchparams::getAllUSERParams for an example of using this method.
| query | should be in the range 1..getNumQueries(). |
| num | is the line number (1..n) of the line in the required section. |
| pPreviousNum | is an optional parameter to speed up iterating through a large section when using cached files. If this value is not null, then the pPreviousOffset should also be passed. |
| pPreviousOffset | is an optional parameter to speed up iterating through a large section when using cached files. If this value is not null, then the pPreviousNum should also be passed. |
| std::string enumerateSectionKeys | ( | const section | sec, |
| const int | num, | ||
| int * | pPreviousNum = 0, |
||
| OFFSET64_T * | pPreviousOffset = 0 |
||
| ) | const |
Get the key name for each item in a section.
Enumerate through all the entries in a section. It returns the key name (not value) for a single item.
In version of Mascot Parser prior to version 2.3, the items are returned in alphabetical order. In Mascot Parser version 2.3 and later, the items are returned in the order in the results file.
See ms_searchparams::getAllUSERParams for an example of using this method.
| sec | can be any section number that contains value= lines This function will not work for sections that contain embedded XML such as the unimod section. |
| num | is the line number (1..n) of the line in the required section. |
| pPreviousNum | is an optional parameter to speed up iterating through a large section when using cached files. If this value is not null, then the pPreviousOffset should also be passed. |
| pPreviousOffset | is an optional parameter to speed up iterating through a large section when using cached files. If this value is not null, then the pPreviousNum should also be passed. |
| std::string get_ms_mascotresults_params | ( | const ms_mascotoptions & | opts, |
| ms_mascotresults_params & | resParams | ||
| ) | const |
Return default flags and parameters for creating an ms_peptidesummary or ms_proteinsummary object.
A number of optional flags and parameters can be passed to the ms_proteinsummary or ms_peptide summary constructors. For an application or script running on the Mascot server, the default values for some of these parameters should normally be taken from the mascot.dat file. This function sets the values and flags required to pass to the constructor in the passed ms_mascotresults_params object.
| [in] | opts | contains the options stored in mascot.dat. Call the ms_datfile construction and then ms_datfile::getMascotOptions() to obtain this value. |
| [out] | resParams | the values and flags required to pass to the peptide or protein summary object are set to this ms_mascotresults_params object, overwriting any values which were already set. |
| std::string get_ms_mascotresults_params | ( | const ms_mascotoptions & | opts, |
| unsigned int * | gpFlags, | ||
| double * | gpMinProbability, | ||
| int * | gpMaxHitsToReport, | ||
| double * | gpIgnoreIonsScoreBelow, | ||
| unsigned int * | gpMinPepLenInPepSummary, | ||
| bool * | gpUsePeptideSummary, | ||
| unsigned int * | gpFlags2 | ||
| ) | const |
[Deprecated] Return default flags and parameters for creating an ms_peptidesummary or ms_proteinsummary object.
A number of optional flags and parameters can be passed to the ms_proteinsummary or ms_peptide summary constructors. For an application or script running on the Mascot server, the default values for some of these parameters should normally be taken from the mascot.dat file. This function returns the values and flags required to pass to the constructor.
See Multiple return values in Perl, Java, Python and C#.
| [in] | opts | contains the options stored in mascot.dat. Call the ms_datfile construction and then ms_datfile::getMascotOptions() to obtain this value. |
| [out] | gpFlags | will return the flags that are to be passed as the second parameter to the ms_proteinsummary or ms_peptidesummary object. |
| [out] | gpMinProbability | is the third parameter to be passed to the ms_proteinsummary or ms_peptidesummary objects. This return value will nomally be equal to the value returned from ms_mascotoptions::getSigThreshold(). |
| [out] | gpMaxHitsToReport | this return value will normally be the one returned by ms_searchparams::getREPORT(). |
| [out] | gpIgnoreIonsScoreBelow | this return value will be the one returned by ms_mascotoptions::getIgnoreIonsScoreBelow(). |
| [out] | gpMinPepLenInPepSummary | this return value will be the one returned by ms_mascotoptions::getMinPepLenInPepSummary. |
| [out] | gpUsePeptideSummary | will be false is the file doesn't contain any anyMSMS (as returned by the anyMSMS() function). In this case, you should create an ms_proteinsummary. If gpUsePeptideSummary is true, you should create an ms_peptidesummary object. |
| [out] | gpFlags2 | is only required for an ms_peptidesummary. If gpUsePeptideSummary is true, gpFlags2 will have the following bits set.
|
| std::string getCacheDirectory | ( | bool | processed = true ) |
const |
Returns the directory being used for cache files (if any).
The cacheDirectory supplied to the constructor ms_mascotresfile::ms_mascotresfile may contain a number of '%' flags which get substituted by Mascot Parser.
This function returns either an absolute directory, or a directory relative to the current working directory.
See Caching Mascot Results and ms_mascotoptions::getCacheDirectory
| processed | if true (the default), then the returned directory is relative to the current directory and will have any '%' flags replaced with the relevant directory. If processed is false, then the directory returned will be identical to the one passed to the constructor. |
| std::string getCacheFileName | ( | ) | const |
Returns the filename of the cache file.
If the cacheDirectory supplied to the ms_mascotresfile::ms_mascotresfile constructor is not empty, then a filename will be returned. This does not guarantee that the file exists, or is being used.
| bool getCrosslinkingMethod | ( | ms_crosslinking_method * | method ) | const |
Return the crosslinking method object from the crosslinking section of the results file.
This method returns true if all of the following apply:
CROSSLINKING parameter exists; Otherwise the method returns false.
If the CROSSLINKING parameter is empty or equals "none", then the method simply returns false. Otherwise, on failure, the method sets the warning ms_errors::ERR_MSP_CROSSLINKING_FAILEDLOAD.
| method | A pointer to crosslinking method object. This must be a valid pointer to a valid object, which should normally be created using the default constructor ms_quant_method::ms_quant_method. |
| DATABASE_TYPE getDatabaseType | ( | const int | idx ) | const |
Return database type if available.
Mascot 2.6 and later save the type of the searched database(s) in the results file, as db_typeX= lines in the header section. These types are AA (amino acid), NA (nucleic acid) or SL (spectral library). The number of databases is ms_searchparams::getNumberOfDatabases(), so idx should be between 1 and getNumberOfDatabases().
Spectral libraries must have a reference database. If the reference database is not part of the actual search, protein accessions mapped to it have a database number above ms_searchparams::getNumberOfDatabases(). For example, if the search contains one AA database and one spectral library, getNumberOfDatabases() is 2 and the types returned by getDatabaseType() are AA (idx = 1) and SL (idx = 2). The reference database is at index 3 with type SLREF.
To find the number of the reference database of a spectral library, see ms_mascotresfile::geReferenceDatabaseNumberOfSL().
| idx | index of the database; must normally be between 1 and ms_searchparams::getNumberOfDatabases(), or a valid database number returned by ms_mascotresfile::getReferenceDatabaseNumberOfSL() |
| int getDate | ( | ) | const [inline] |
Returns the date and time of the search in seconds since midnight January 1st 1970.
Obtained from the date= line in the header section of the file. Can be converted to day, month, year etc. using gmtime or similar functions.
| ms_mascotoptions::DECOY_ALGORITHM getDecoyTypeForDB | ( | const int | idx = 1 ) |
const |
Returns the decoy algorithm type for a given database.
Decoy algorithm type is saved as decoy_type= or decoy_typeX= in the header section, depending on Mascot version.
If idx = 1, the method returns the value of decoy_type=. If idx > 1, the method returns the corresponding decoy_typeX= line, or if one doesn't exist, falls back on decoy_type=.
If there is no suitable value in the file or idx is outside its range, the method returns ms_mascotoptions::DECOY_ALGORITHM_NONE.
| idx | index of the database from 1 to ms_searchparams::getNumberOfDatabases(). |
| bool getEnzyme | ( | ms_enzymefile * | efile, |
| const char * | enzymeFileName = 0 |
||
| ) | const |
Returns an object that represents enzyme-section as a reduced enzymes file.
For data files created with Mascot 2.2 and later, the full definition of the enzyme used is included in the Mascot results file. For earlier versions of Mascot, just the name is recorded. This function attempts to read the definition from the results file. If the definition is not present in the results file and a path to the enzymes file has been passed, then this function reads the enzymes file and removes all entries from the list in memory apart from the one with the name specified in the results file.
To determine whether the content has been parsed successfully call ms_enzymefile::isValid.
See Object initialising functions in Perl, Java, Python and C#.
| efile | a pointer to enzymes-file object that will accept the content from the section or the enzymes file if necessary. If successful, the enzyme itself can be retrieved by passing an index of zero to ms_enzymefile::getEnzymeByNumber() |
| enzymeFileName | is only used for results files prior to Mascot 2.2 |
| const ms_errs * getErrorHandler | ( | ) | const [inherited] |
Retrive the error object using this function to get access to all errors and error parameters.
See Error Handling.
| int getErrorNumber | ( | const int | num = -1 ) |
const |
Return a specific error number - or ms_mascotresfile::ERR_NO_ERROR.
All errors are accumulated into a list in this object, until clearAllErrors() is called.
Errors in other classes are accumulated here. If, for example, there is an error when creating a peptide summary, the errors need to be accessed through this class.
See Error Handling.
In Mascot Parser 2.5 and later, this is implemented by calling: ms_errs::getErrorNumber()
| num | is the error number in the range 1..getNumberOfErrors(). Passing a value of -1 will return the last error, or ERR_NO_ERROR. If an invalid number is passed, ERR_NO_ERROR will be returned (and no error will be added to the list of errors!). |
| std::string getErrorString | ( | const int | num ) | const |
Return a specific error as a string.
All errors are accumulated into a list in this object, until clearAllErrors() is called. To return a particular error, call this function with a number 1..getNumberOfErrors(). Passing a value of -1 will return the last error, or an empty string. If an invalid number is passed an empty string will be returned (and no error will be added to the list of errors!).
Errors in other classes are accumulated here. If, for example, there is an error when creating a peptide summary, the errors need to be accessed through this class.
In Mascot Parser 2.5 and later, this is implemented by calling ms_errs::getErrorString but functionality is identical to previous versions.
See Error Handling.
| num | 1 to number of errors, or -1 |
| int getExecTime | ( | ) | const [inline] |
Returns the time taken for the search.
Obtained from the exec_time= line in the header section. This is the 'wall clock' time, not the CPU time.
| std::string getFastaPath | ( | int | idx = 1 ) |
const |
Returns the path to the FASTA file used.
Available in Mascot 2.2 and later.
Mascot 2.6 and later support spectral libraries. Each spectral library must have a reference database into which found peptide sequences are mapped at the end of the search. The "effective" reference database could be one of the protein sequence databases searched, or it could be a separate database used only for lookup purposes. You can find the database number of the reference database with ms_mascotresfile::getReferenceDatabaseNumberOfSL(). If this is larger than getNumberOfDatabases(), the FASTA file path is obtained from the library_reference_fastafile line in the header section of the results file.
| idx | index of the database from 1 to ms_searchparams::getNumberOfDatabases() for databases or libraries searched, or an index returned by ms_mascotresfile::getReferenceDatabaseNumberOfSL(). |
| std::string getFileName | ( | const int | id = 1 ) |
const |
Returns the name of the results file passed into the constructor.
| id | a 1 based index. Unless appendResfile() has been called, this value must be '1'. |
| int getJobNumber | ( | const int | resfileID = 1 ) |
const |
Return the job number for this file - obtained from the file name.
The library can only 'guess' at this since the value is not in the results file. To perform this function, it retrieves the job number from the file name, so be warned about changing file names. The function returns 0 if it cannot determine the job number.
| resfileID | is the the 1 based id of the results file. If multiple files have been merged together with appendResfile(), use the file ID returned by appendResfile() or getSrcFileIdForMultiFile() to access the job number of the appended files. |
| void getKeepAlive | ( | KA_TASK & | kaTask, |
| int & | kaPercentage, | ||
| std::string & | kaAccession, | ||
| int & | kaHit, | ||
| int & | kaQuery, | ||
| std::string & | kaText | ||
| ) | const |
Return the progress indicators used by the keepAlive functions.
See Multiple return values in Perl, Java, Python and C# although there may be issues with some languages and the kaTask parameter.
It is normally easier for client applications to call ms_mascotresults::getCreateSummaryProgress() or outputKeepAlive() than to call this function.
| kaTask | is the task currently being performed by Mascot Parser. If Parser is doing nothing, then this will be the last task that was completed and kaPercentage will be 100 |
| kaPercentage | is the percentage (0..100) complete for the current kaTask. |
| kaAccession | is the current 'accession' being processed. See outputKeepAlive() for details of which tasks set this value |
| kaHit | is the current hit being processed. See outputKeepAlive() for details of which tasks set this value |
| kaQuery | is the current 'query' being processed. See outputKeepAlive() for details of which tasks set this value |
| kaText | is the text that would be output by outputKeepAlive() |
| int getLastError | ( | ) | const |
Return the last error number - or ms_mascotresfile::ERR_NO_ERROR.
Same as calling getErrorNumber() with -1 as a parameter.
Reimplemented from ms_errors.
| std::string getLastErrorString | ( | ) | const |
Return the last error number - or an empty string.
Same as calling getErrorString() with -1 as a parameter.
Reimplemented from ms_errors.
| std::string getMascotVer | ( | ) | const [inline] |
Returns the version of Mascot used to perform the search.
Obtained from the version= entry in the header section of the file.
| bool getMasses | ( | ms_masses * | masses ) | const |
Returns an ms_masses object from the mass values in the results file.
For results files from Mascot 2.2 and later, this function simply calls
if (getUnimod(&umodConfigFile) && umodConfigFile.isValid())
{
masses->copyFrom(&umodConfigFile);
}
For earlier results files, it reads the residue mass values from the masses section of the file. Since this section either contains average or monoisotopic masses (but not both), the resulting ms_masses object will only have one set of masses.
See Object initialising functions in Perl, Java, Python and C#.
| masses | a pointer to a valid masses object that will accept the content from the section. |
| const ms_modification * getMonoLinkModification | ( | const int | modNum, |
| const int | monoLink | ||
| ) | const |
Returns an ms_modification object that represents a monolink variable modification.
The method performs the following steps:
The ms_modification object contains the following fields:
| modNum | Variable mod index in range 1..32. Usually this comes from the peptide's variable mods string. |
| monoLink | Index of the neutral loss element. Usually this comes from the peptide's monolink string. |
| std::string getMSParserVersion | ( | ) | const |
Returns the version number of the Mascot Parser library.
This version information is also available:
perl -Mmsparser -e "print msparser->VERSION()"
JarFile jar = new JarFile(new File(jarName));
Manifest jarManifest = jar.getManifest();
Attributes mainAttributes = jarManifest.getMainAttributes();
String version = (String) mainAttributes.get(Attributes.Name.IMPLEMENTATION_VERSION);
| int getMultiFileQueryNumber | ( | const int | localQuery, |
| const int | fileId | ||
| ) | const |
Return the multi-file query number from the local query number in an appended file.
See Multiple return values in Perl, Java, Python and C#.
Needs to be called on the 'primary' file object rather than on a ms_mascotresfile object returned by the getResfile() function.
See Combining multiple .dat files.
Example: Assume that the primary .dat file as 10 queries, file 2 has 20 queries and file 3 has 30 queries. This function will return the following:
| localQuery | fileId | returned query |
|---|---|---|
| 6 | 1 | 6 |
| 11 | 2 | 21 |
| 1 | 3 | 31 |
See also: getSrcQueryAndFileIdForMultiFile() for the 'inverse' function
| localQuery | is the query number which should be a value between 1 and getNumQueries() for the ms_mascotresfile specified by fileId. |
| fileId | is a 1 based index to the source .dat file. |
| int getNumberOfErrors | ( | ) | const |
Return the number of errors since the last call to clearAllErrors.
This will be zero if there has been no error.
All errors are accumulated into a list in this object, until clearAllErrors() is called.
Errors in other classes are accumulated here. If, for example, there is an error when creating a peptide summary, the errors need to be accessed through this class.
From version 2.5, implemented by calling getErrorHandler()->getNumberOfErrors()
See Error Handling.
| int getNumberOfResfiles | ( | ) | const |
Multiple results files can be summed together and treated as 'one'.
See Combining multiple .dat files.
| int getNumEtSeqsSearched | ( | const int | idx = 0 ) |
const |
Returns the number of sequences searched in the second pass of an integrated error tolerant search.
Obtained from the et_sequences= or et_sequencesX= line in the header section of the file.
See Integrated error tolerant search . This function will return -1 for the Original error tolerant search and for searches prior to Mascot 2.4.1
| idx | index of the database. Specifying a value of 0 (the default) will return the sum of all sequences after taxonomy in all FASTA files searched. Otherwise, the value should be in the range from 1 to ms_searchparams::getNumberOfDatabases(). Specifying a value outside the range of 0..ms_searchparams::getNumberOfDatabases() will result in -1 being returned. If the database at index idx is a spectral library, the value returned is -1. |
| int getNumHits | ( | const section | sec = SEC_SUMMARY ) |
const |
Returns the maximum number of hits possible for a protein summary.
If no parameter is passed to this function, information is returned about the standard 'summary' section.
This function returns the number of hits contained in the summary, decoy_summary, et_summary or et_decoy_summary section of the results file. The value returned by ms_proteinsummary::getNumberOfHits will generally be more useful. The number of hits recorded in the summary section sec could be zero even if there are peptide matches in the corresponding peptides section.
If appendResfile() has been called (see Combining multiple .dat files) then this value will return -1 because the protein summary is not supported for multiple dat files.
| sec | Can be one of ms_mascotresfile::SEC_SUMMARY, ms_mascotresfile::SEC_DECOYSUMMARY, ms_mascotresfile::SEC_ERRTOLSUMMARY, ms_mascotresfile::SEC_ERRTOLDECOYSUMMARY. |
num_hits= in the specified section or -1 if an invalid section is passed of if the object is a multifile results file. | int getNumLibraryEntries | ( | const int | idx = 0 ) |
const |
Returns the number of entries in the spectral library searched.
Obtained from the library_entriesX= line in the header section of the file. Spectral library support was added in Mascot 2.6.
| idx | index of the database. Specifying a value of 0 (the default) will return the sum of all library entries searched. Otherwise, the value should be in the range from 1 to ms_searchparams::getNumberOfDatabases(). Specifying a value outside the range of 0..ms_searchparams::getNumberOfDatabases() will result in -1 being returned. If the database at index idx is not a spectral library, the value returned is -1. |
| int getNumQueries | ( | const int | resfileID = 0 ) |
const |
Returns the number of queries (peptide masses or ms-ms spectra).
Obtained from the queries= line in the header section of the file.
| resfileID | is the the 1 based id of the results file. When the default value of 0 is used for a single .dat file, this is number of queries in the file. For Combining multiple .dat files supplying a value of zero returns the total number of queries in all the results files that have been combined. Use a value of 1 to get the number of queries just in the first .dat file. |
| double getNumResidues | ( | const int | idx = 0 ) |
const |
Returns the number of residues in the FASTA file(s) searched.
Obtained from the residues= or residuesX= line in the header section of the file. Multiple FASTA file support was added in Mascot 2.3.
| idx | index of the database. Specifying a value of 0 (the default) will return the sum of all sequences after taxonomy in all FASTA files searched. Otherwise, the value should be in the range from 1 to ms_searchparams::getNumberOfDatabases(). Specifying a value outside the range of 0..ms_searchparams::getNumberOfDatabases() will result in -1 being returned. If the database at index idx is a spectral library, the value returned is 0. |
| int getNumSeqs | ( | const int | idx = 0 ) |
const |
Returns the number of sequences in the FASTA file(s) searched.
Obtained from the sequences= or sequencesX= in the header section of the file. Multiple FASTA file support was added in Mascot 2.3
| idx | index of the database. Specifying a value of 0 (the default) will return the sum of all sequences in all FASTA files searched. Otherwise, the value should be in the range from 1 to ms_searchparams::getNumberOfDatabases(). Specifying a value outside the range of 0..ms_searchparams::getNumberOfDatabases() will result in -1 being returned. If the database at index idx is a spectral library, the value returned is 0. |
| int getNumSeqsAfterTax | ( | const int | idx = 0 ) |
const |
Returns the number of sequences that passed the taxonomy filter in the FASTA file(s) searched.
Obtained from the sequences_after_tax= or sequences_after_taxX= line in the header section of the file. Multiple FASTA file support was added in Mascot 2.3.
| idx | index of the database. Specifying a value of 0 (the default) will return the sum of all sequences after taxonomy in all FASTA files searched. Otherwise, the value should be in the range from 1 to ms_searchparams::getNumberOfDatabases(). Specifying a value outside the range of 0..ms_searchparams::getNumberOfDatabases() will result in -1 being returned. If the database at index idx is a spectral library, the value returned is 0. |
| int getObservedCharge | ( | const int | query, |
| const bool | decoy = false |
||
| ) | const |
The 'charge' returned will be 0 for Mr, otherwise it will be 1, -1, 2, -2, 3, -3 etc. and -100 for an error.
This is obtained from the qexp[query] value. It will come from the SEC_SUMMARY section unless decoy is set to true in which case it is obtained from the SEC_DECOYSUMMARY section.
The 'charge' returned will be 0 for Mr, otherwise it will be 1, -1, 2, -2, 3, -3 etc. and -100 for an error.
Possible error values:
If an 'ambiguous' charge state is specified for the whole search or for a specific query, then Mascot just records matches for the highest scoring charge state, and it is this charge state that is returned from this function. For example, the search may have been performed with "2+, 3+ or 4+" and ms_inputquery::getCharge() will return "2+, 3+ or 4+". If the highest scoring peptide match for a particular query was to charge state 4+, then all top 10 matches for that query will be for 4+ and this function will return '4'.
It is therefore not impossible to get different charge values from the decoy and standard sections of the results files.
This method can only read charge from the 'summary' and 'decoy_summary' sections. If the search is error tolerant or intact crosslink search, the 'et_summary' and 'crosslink_summary' sections can report charge states different from the 'summary' section. It's best to use ms_peptide::getCharge().
The functions getObservedIntensity() and getObservedMass() do not require the decoy parameter as the values will be identical from the SEC_SUMMARY and the SEC_DECOYSUMMARY sections.
| query | is a number in the range 1..getNumQueries() |
| decoy | should be false for SEC_SUMMARY and true for SEC_DECOYSUMMARY. |
| double getObservedIntensity | ( | const int | query ) | const |
Returns the experimental intensity for the peptide.
This is obtained from SEC_SUMMARY - qintensity[query]. This value is not always available and does not need to be supplied by the end user.
Returns 0 if the query cannot be found and sets the error ms_mascotresfile::ERR_QUERYOUTOFRANGE.
| query | is a number in the range 1..getNumQueries() |
| double getObservedMass | ( | const int | query ) | const |
Returns the experimental mass value as entered by the user.
This is obtained from SEC_SUMMARY - qexp[query].
Returns 0 if the value cannot be found and sets the error ms_mascotresfile::ERR_QUERYOUTOFRANGE.
| query | query number |
| double getObservedMrValue | ( | const int | query, |
| const bool | decoy = false |
||
| ) | const |
Returns the experimental mass value (as a relative mass) as entered by the user.
This is obtained from the qmass[query] value. It will come from the SEC_SUMMARY section unless decoy is set to true in which case it is obtained from the SEC_DECOYSUMMARY section.
Returns 0 if the value cannot be found and sets the error ms_mascotresfile::ERR_QUERYOUTOFRANGE.
| query | is a number in the range 1..getNumQueries() |
| decoy | should only be set to true if ms_searchparams::getDECOY returns true. |
| std::vector< std::string > getPercolatorFileNames | ( | ) | const |
Retrieve the filenames use for percolator input and output.
This function will return an empty vector unless setPercolatorFeatures() has been called before calling this.
The offsets into the return array are defined by ms_mascotresfile::PERCOLATOR_FILE_NAMES.
See Using STL vector classes vectori, vectord and VectorString in Perl, Java, Python and C#.
| ms_progress_info * getProgressInfo | ( | bool | forPeptideSummary = false ) |
const |
If a matrix_science::ms_progress_info object is passed to the constructor, this is returned here.
| forPeptideSummary | should be set to true to get the progress object used when creating the ms_peptidesummary, or false to return the object used when creating the ms_mascotresfile |
| bool getQuantitation | ( | ms_quant_configfile * | qfile ) | const |
Returns an object that represents the quantitation section as a reduced quantitation.xml file.
The return value indicates that the section exists. Call ms_quant_configfile::isValid to determine whether the XML part has been parsed successfully.
For quantitation_2 and later, the contents of the file are validated against a schema by default. For quantitation_1, to explicitly validate against a schema, use ms_quant_configfile::setSchemaFileName() to choose a schema, and then use ms_quant_configfile::validateDocument() to validate.
See Object initialising functions in Perl, Java, Python and C#.
| qfile | a pointer to quantitation file object. This must be a valid pointer to a valid object, which should normally be created using the default constructor: ms_quant_configfile::ms_quant_configfile |
| bool getQuantitationMethod | ( | ms_quant_method * | qm ) | const |
Return the quantitation method object from the quantitation section of the results file.
This method returns true if all of the following apply:
QUANTITATION parameter exists; Otherwise the method returns false.
If the QUANTITATION parameter is empty or equals "none", then the method simply returns false. Otherwise, on failure, the method sets the warning ms_errors::ERR_MSP_QUANT_FAILEDLOAD.
| qm | A pointer to quantitation method object. This must be a valid pointer to a valid object, which should normally be created using the default constructor ms_quant_method::ms_quant_method. |
| int getQuerySectionValue | ( | const int | query, |
| const char * | key, | ||
| char * | str, | ||
| int | maxLen | ||
| ) | const |
Return the string value from a query in the results file.
Returns a string from the query section. It is generally easier to use the ms_inputquery class rather than use this lower level function.
| query | is a number in the range 1..getNumQueries() |
| key | is case insensitive. |
| str | is a pointer to a buffer to receive the string. |
| maxLen | is the length of the buffer (including a null terminator). |
| double getQuerySectionValueDouble | ( | const int | query, |
| const char * | key | ||
| ) | const |
Return the floating point value from a query in the results file.
Gets the number from the query section of the results file if it exists.
| query | is a number in the range 1..getNumQueries() |
| key | is case insensitive. |
| int getQuerySectionValueInt | ( | const int | query, |
| const char * | key | ||
| ) | const |
Return the integer value from a query in the results file.
Gets the number from the query section of the results file if it exists.
| query | is a number in the range 1..getNumQueries() |
| key | is case insensitive. |
| std::string getQuerySectionValueStr | ( | const int | query, |
| const char * | key | ||
| ) | const |
Return the string value from a query in the results file.
Gets the string from the query section of the results file if it exists. It is generally easier to use the ms_inputquery class rather than use this lower level function.
| query | is a number in the range 1..getNumQueries() |
| key | is case insensitive. |
| int getReferenceDatabaseNumberOfSL | ( | const int | idx ) | const |
Return the database number of the reference database of a spectral library.
The reference database of a spectral library is either one of the databases searched -- if the reference database was part of the actual search -- or a virtual database whose number is above ms_searchparams::getNumberOfDatabases(). In the first case, getReferenceDatabaseNumberOfSL() returns a database number between 1 and ms_searchparams::getNumberOfDatabases(). In the second case, the number is above getNumberOfDatabases().
If idx does not refer to a spectral library, the method returns -1.
| idx | Database number of the spectral library of interest, between 1 and ms_searchparams::getNumberOfDatabases() and whose type is SL (see getDatabaseType()) |
| std::string getRepeatSearchString | ( | const int | query, |
| const bool | fullQuery = false |
||
| ) | const |
To perform a repeat search need to build up appropriate string.
If the 'fullQuery' parameter is false (the default) then the format will be:
mr from(observed, charge) query(querynum) etc.
If the 'fullQuery' parameter is true then the format will be:
mr from(observer, charge1, charge2...) ions() etc.
where 'etc' will be one or more of the following (split onto several lines here for readability):
intensity(value) peptol(value, units) seq() comp() tag() etag() title() instrument() it_mods() index() rtinseconds() rtinseconds[x]() scans scans[x]() rawscans rawscans[x]()
mr value is for the first charge. PepTol= line in the queryx section of the results file. seq1= line in the queryx section of the results file. If there were multiple seq() commands in the original search, then these will be in the results file as seq1=, seq2= etc., and will be returned as seq(...) seq(...). comp1= line in the queryx section of the results file. If there were multiple comp() commands in the original search, then these will be in the results file as comp1=, comp2= etc., and will be returned as comp(...) comp(...). tag1=t line in the queryx section of the results file. If there were multiple tag() commands in the original search, then these will be in the results file as tag1=, tag2= etc., and will be returned as tag(...) tag(...). tag1=e line in the queryx section of the results file. If there were multiple etag() commands in the original search, then these will be in the results file as tag1=, tag2= etc., and will be returned as etag(...) etag(...). scans(29-34, 43) or scans[0](29-34) scans[1](43). rtinseconds(10-20, 25) or rtinseconds[0](10-20) rtinseconds[1](25). rawscans[0](pd0cy1ex1:pd0cy1ex3) rawscans[1](fn2ix1). rawfile(c:/data/rawfile.raw). locus(2.1.1.24.1). nph-mascot.exe comes to the query(x) command it gets the ions vales from the original .dat file Returns 0 if the value cannot be found and sets the error ms_mascotresfile::ERR_QUERYOUTOFRANGE.
See Automated repeating of searches.
| query | is a number in the range 1..getNumQueries() |
| fullQuery | If true, then a complete and self contained sequence query will be returned. See above for details |
| const ms_mascotresfile * getResfile | ( | int | resfileID ) | const |
Returns a pointer to the resfile created by calling appendResfile.
See Combining multiple .dat files and Maintaining object references: two rules of thumb.
| resfileID | is the the 1 based id of the results file and must be in the range 1..getNumberOfResfiles(). A value of 1 will (not particularly usefully!) return a pointer to the ms_mascotresfile that was originally created. A value of 2 will return ms_mascotresfile created in the first successful call to appendResfile(), and so on. |
| const char * getSectionName | ( | const section | sec ) | [static] |
Return the section name as a string given the enum value.
The section name is intended for printing in error or log messages.
| sec | One of the values in ms_mascotresfile::section. |
| int getSectionValue | ( | const section | sec, |
| const char * | key, | ||
| char * | str, | ||
| int | maxLen | ||
| ) | const |
Return the string value from any line in the results file.
Function to return a string (into the passed str value) from the results file without using STL. It is generally easier to use getSectionValueStr() which returns a std::string.
Gets the requested string from the results file if it exists. Use the enumeration to choose which section. Returns the number of characters copied into 'str' which will not be greater than maxLen. E.g.
char s[1024]; f.getSectionValue(ms_mascotresfile::SEC_PARAMETERS,"MODS",s,1024);
| sec | can be any section number |
| key | is case insensitive. |
| str | is a pointer to a buffer to receive the string |
| maxLen | is the length of the buffer (including a null terminator) |
| double getSectionValueDouble | ( | const section | sec, |
| const char * | key | ||
| ) | const |
Return the floating point value from any line in the results file.
| sec | can be any section number |
| key | is case insensitive. |
| int getSectionValueInt | ( | const section | sec, |
| const char * | key | ||
| ) | const |
Return the integer value from any line in the results file.
A value of -1 will be returned if there is no corresponding entry in the results file.
A value of zero will be returned if the entry contains a value that isn't a number.
If the correct value is out of the range of representable values, INT_MAX or INT_MIN is returned.
| sec | can be any section number |
| key | is case insensitive. |
| std::string getSectionValueStr | ( | const section | sec, |
| const char * | key | ||
| ) | const |
Return the string value from any line in the results file.
Gets the requested string from the results file if it exists. Use the enumeration to choose which section. E.g.
std::string s;
s = file.getSectionValue(ms_mascotresfile::SEC_PARAMETERS, "MODS");
| sec | can be any section number |
| key | is case insensitive. |
| std::vector< int > getSLDatabaseNumbersOfReference | ( | const int | idx ) | const |
Return the database numbers of the spectral libraries whose reference database is at the given index.
The reference database of a spectral library is either one of the databases searched -- if the reference database was part of the actual search -- or a virtual database whose number is above ms_searchparams::getNumberOfDatabases(). The same database can be the reference database of multiple spectral libraries.
If idx does not refer to a reference database, the method returns the empty vector.
| idx | Database number of the reference database of interest. |
| double getSLFragmentTolerance | ( | int | idx = 1 ) |
const |
Returns the effective spectral library fragment tolerance.
When a search is run against a spectral library, the effective fragment tolerance is calculated from the user-configured library fragment tolerance and the tolerance specified in the search parameters. This may be different from ms_searchparams::getITOL() and ms_searchparams::getITOLU().
The tolerance unit can be retrieved with getSLFragmentToleranceUnit().
In Mascot 2.6.01 and later, the effective tolerance value and unit are saved in the header section of the results file as sl_itolX=. For files created by Mascot 2.6.00, the value is parsed from the sl_exec_commandX= line if present.
If idx is outside its range, the method returns 0.0.
| idx | index of the database from 1 to ms_searchparams::getNumberOfDatabases(). |
| std::string getSLFragmentToleranceUnit | ( | int | idx = 1 ) |
const |
Returns the unit of the effective spectral library fragment tolerance.
When a search is run against a spectral library, the effective fragment tolerance is calculated from the user-configured library fragment tolerance and the tolerance specified in the search parameters. This may be different from ms_searchparams::getITOL() and ms_searchparams::getITOLU().
The tolerance can be retrieved with getSLFragmentTolerance().
In Mascot 2.6.01 and later, the effective tolerance value and unit are saved in the header section of the results file as sl_itolX=. For files created by Mascot 2.6.00, the value is parsed from the sl_exec_commandX= line if present.
If idx is outside its range, the method returns 0.0.
| idx | index of the database from 1 to ms_searchparams::getNumberOfDatabases(). |
| bool getSrcQueryAndFileIdForMultiFile | ( | const int | q, |
| int & | gsqNewQuery, | ||
| int & | gsqFileId | ||
| ) | const |
Return the query number and file ID in the source .dat file.
See Multiple return values in Perl, Java, Python and C#.
Useful for combining multiple results files (see Combining multiple .dat files) but also returns valid values for a single file.
Example: Assume that the primary .dat file as 10 queries, file 2 has 20 queries and file 3 has 30 queries. This function will return the following:
| q | newQuery | fileId |
|---|---|---|
| 6 | 6 | 1 |
| 21 | 11 | 2 |
| 31 | 1 | 3 |
See also: getMultiFileQueryNumber() for the 'inverse' function
| q | is the query number which should be a value between 1 and getNumQueries(). |
| gsqNewQuery | is used to return the query number in the specified source file. |
| gsqFileId | is a 1 based index to the source .dat file. |
| bool getTaxonomy | ( | ms_taxonomyfile * | tfile ) | const |
Returns an object that represents taxonomy-section as a reduced taxonomy file.
The return value only indicates that the section exists. If you want to find out whether the content has been parsed successfully, call the methods of matrix_science::ms_taxonomyfile.
See Object initialising functions in Perl, Java, Python and C#.
| tfile | a pointer to taxonomy file object that will accept the content from the section. |
| bool getUnimod | ( | ms_umod_configfile * | ufile, |
| bool | useSchemaFromResfile = false |
||
| ) | const |
Returns an object that represents unimod-section as a reduced unimod_2.xml file.
The return value only indicates that the section exists. If you want to find out whether the XML part has been parsed successfully, call ms_umod_configfile::isValid
See Object initialising functions in Perl, Java, Python and C#.
| ufile | a pointer to unimod file object that will accept the content from the section. |
| useSchemaFromResfile | determines where the location of the xml schema is defined. If 'true', then the schema location should have been defined by specifying XMLschemaDirectory in the constructor. If 'false', then ms_umod_configfile::setSchemaFileName must have been called on ufile before calling this function. This parameter was added in Parser 2.5, and the default value is false to ensure that it is backward compatible with previous versions. |
| bool getUnimodXL | ( | ms_umod_configfile * | ufile, |
| bool | useSchemaFromResfile = false |
||
| ) | const |
Returns an object that represents unimod_xl-section as a reduced unimod_xl.xml file.
The return value only indicates that the section exists. If you want to find out whether the XML part has been parsed successfully, call ms_umod_configfile::isValid
See Object initialising functions in Perl, Java, Python and C#.
| ufile | a pointer to unimod file object that will accept the content from the section. |
| useSchemaFromResfile | determines where the location of the xml schema is defined. If 'true', then the schema location should have been defined by specifying XMLschemaDirectory in the constructor. If 'false', then ms_umod_configfile::setSchemaFileName must have been called on ufile before calling this function. ms_mascotresfile::getUnimodXL uses the same schema as ms_mascotresfile::getUnimod. |
| std::string getUniqueTaskID | ( | ) | const |
Returns the unique task ID used by Mascot Daemon.
Although this value is a number, it is a 64 bit integer. Some languages on some platforms cannot deal with 64 bit integers properly, so the value is returned as a string. For searches with no taskid, an empty string is returned.
| std::string getXMLschemaFilePath | ( | XML_SCHEMA | XMLschema ) | const |
Gets the XML schema to be used by functions using quantitation or unimod.
| XMLschema | The XML_SCHEMA enumeration value of the required xml schema file. |
| bool isDatabaseTypeAvailable | ( | ) | const |
Check whether database types are available.
Mascot 2.6 and later save the type of the database(s) in the results file, as db_typeX= lines in the header section. If the types are not available, the database or databases searched could be AA or NA.
| bool isErrorTolerant | ( | ) | const |
Returns true if the search was an error tolerant search.
Obtained from the ERRORTOLERANT value in the parameters section. In Mascot versions 1.8 and later, an error tolerant search can be run as a repeat search. In this case, one or more ACCESSIONs (which may be retrieved using ms_searchparams::getACCESSION ) must have been specified, and the results file will just contain the error tolerant search results. In Mascot 2.2 and later, a single search can be performed which contains both the standard search results and the error tolerant search results of automatically selected proteins. In this case, ms_searchparams::getACCESSION will return an empty string.
| bool isMSMS | ( | ) | const |
Returns true if the search was an MSMS search (SEARCH=MIS).
Since all types of search may be entered as a sequence query, it may be more useful to use the anyMSMS() member.
SEARCH=MIS in the parameters section of the file. | bool isPMF | ( | ) | const |
Returns true if the search was a PMF search (SEARCH=PMF).
Since all types of search may be entered as a sequence query, it may be more useful to use the anyPMF() member.
SEARCH=PMF in the parameters section of the file. | bool isSQ | ( | ) | const |
Returns true if the search was a sequence query search (SEARCH=SQ).
Since all types of search may be entered as a sequence query, it may be more useful to use the anySQ() member.
SEARCH=SQ in the parameters section of the file. | bool isValid | ( | ) | const [inherited] |
Call this function to determine if there have been any errors.
This will return true unless there have been any fatal errors.
See Error Handling.
| bool outputKeepAlive | ( | ) | const |
Outputs the "keep-alive" string during time-consuming operations.
For HTML reports with large result files it is sometimes necessary to output HTML comments to keep the connection alive. This can be done by specifying the interval at which the text is output (keepAliveInterval) and the text that should be output (keepAliveText) as parameters to the ms_mascotresfile::ms_mascotresfile() constructor.
If the value of keepAliveInterval passed to ms_mascotresfile::ms_mascotresfile is not 0, then the text specfied by keepAliveText will be output approximately every keepAliveInterval seconds. A 'd' in the keepAliveText will be replaced by the number of seconds since the process started.
This functionality is implemented by calling this function 'often', rather than by using a separate thread. This means that the times between calls will not be accurate. A computationally intensive application that uses Mascot Parser can also call this function as required.
From version 2.3 onwards, the keepAliveText can contain tags that allow different text to be output for different lengthy tasks. The tags are:
The text can also include the following tags which are substituted by values:
The following table indicates which values are available for which tasks:
| ci= | rf= | ap= | gp= | ul= | cc= | fd= | |
|---|---|---|---|---|---|---|---|
| %d | X | X | X | X | X | X | X |
| %p | X | X | X | X | X | X | X |
| %h | X | X | X | ||||
| %q | X | X | X | X | |||
| %a | X | X | X | ||||
| %f | X | X | X | X | X | X |
The %a and %h values for cc= are not output for the second half of caching.
A 'complete' example string might be: ' ul=Creating unassigned list (%p% complete)\n qu=Calculating quantitation component intensities (%p% complete)\ ci=Creating cache file (%p% complete)\n rf=Reading results file (%p% complete)\n ap=Assigning peptides to proteins (%p% complete) hit=%h, time=%d\n gp=Found protein group: %a, hit=%h, %p% complete, %d seconds\n cc=Caching results (%p% complete)\n fd=Calculating false discovery rate (%p% complete)\n '
Any text before the first tag will be used as a default for cases where text isn't supplied for a particular task. For example:
Processing: %p% complete gp=Grouping %a
would output the text:
Processing 23% complete
for all tasks except the protein grouping which would output:
Grouping gi|12345
| ms_searchparams& params | ( | ) | const [inline] |
Returns a reference to the search parameters class.
For C# only, params is a keyword, so this function is renamed to _params.
| void resetKeepAlive | ( | const int | keepAliveInterval, |
| const char * | keepAliveText, | ||
| const bool | propagateToAppended = true, |
||
| const bool | resetStartTime = false |
||
| ) |
Replace the existing keepAlive values with new values.
KeepAlive values are passed when creating the ms_mascotresfile object, but it can be useful to change these at a later time.
See outputKeepAlive() for further details.
| keepAliveInterval | is the new interval in seconds. Specify a value of -1 to keep the old value, or 0 to stop outputting keepAliveText. |
| keepAliveText | is the text to output every keepAliveInterval seconds while the file is being loaded, or while other tasks are in progress. |
| propagateToAppended | only has meaning if additional files have been appended by calling appendResfile(). |
| resetStartTime | can be set to true to reset the "%d" value to zero. See outputKeepAlive() for details. |
| void setPercolatorFeatures | ( | const ms_mascotoptions & | options, |
| const char * | additionalFeatures | ||
| ) |
Set Percolator features before creating an ms_peptidesummary with Percolator scoring.
The names (and contents!) of the percolator files depend on the features that have been enabled. This means that, say, turning on an additional feature will cause a new file to be created, but the old file will still be available if the feature is removed again
| options | contains the options stored in mascot.dat It is used to access the percolator features |
| additionalFeatures | is normally a string passed to ms-createpip.exe For example, "-a numUniqPeps -r varmods" would add numUniqPeps and remove varmods from the default |
| void setPercolatorFeatures | ( | const char * | percolatorFeatures, |
| const char * | additionalFeatures, | ||
| const bool | useRetentionTimes | ||
| ) |
Set Percolator features before creating an ms_peptidesummary with Percolator scoring (deprecated).
The names (and contents!) of the percolator files depend on the features that have been enabled. This means that, say, turning on an additional feature will cause a new file to be created, but the old file will still be available if the feature is removed again
| percolatorFeatures | is normally retrieved by calling ms_mascotoptions::getPercolatorFeatures(). |
| additionalFeatures | is normally a string passed to ms-createpip.exe For example, "-a numUniqPeps -r varmods" would add numUniqPeps and remove varmods from the default |
| useRetentionTimes | is a flag to indicate whether retention time information is used by percolator.exe. This value is normally retrieved by calling ms_mascotoptions::isPercolatorUseRT() |
| bool setXMLschemaFilePath | ( | XML_SCHEMA | XMLschema, |
| const char * | path | ||
| ) |
Sets the XML schema to be used by functions using quantitation or unimod.
It is generally easier to pass a directory as the XMLschemaDirectory parameter to the constructor rather than calling this function for each of the required schema.
Example:
std::string qs; qs = "http://www.matrixscience.com/xmlns/schema/quantitation_1 "; qs += "C:/myfiles/quant_schema_1.xsd "; qs += "http://www.matrixscience.com/xmlns/schema/quantitation_2 "; qs += "../schema%20files/quantitation_2.xsd"; setXMLschemaFilePath(XML_SCHEMA_QUANTITATION, qs.c_str()); *
The default values used in cases where this function has not been called and no parameter is passed to the constructor are the values suitable for scripts and programs running on the Mascot Server. These values are:
XML_SCHEMA_QUANTITATION :
http://www.matrixscience.com/xmlns/schema/quantitation_1 ../html/xmlns/schema/quantitation_1/quantitation_1.xsd
http://www.matrixscience.com/xmlns/schema/quantitation_2 ../html/xmlns/schema/quantitation_2/quantitation_2.xsd
XML_SCHEMA_UNIMOD :
http://www.unimod.org/xmlns/schema/unimod_2 ../html/xmlns/schema/unimod_2/unimod_2.xsd
XML_SCHEMA_CROSSLINKING :
http://www.matrixscience.com/xmlns/schema/crosslinking_1 ../html/xmlns/schema/crosslinking_1/crosslinking_1.xsd
| XMLschema | must be one of the valid XML_SCHEMA values |
| path | should be a list of pairs "_schema_alias_ SPACE _file_path_", where SPACE is the space character. See XML_SCHEMA for the supported _scheama_alias_ values for each type of schema |
| bool staticGetPercolatorFileNames | ( | const char * | szDatFileName, |
| const char * | cacheDirectory, | ||
| const char * | percolatorFeatures, | ||
| const char * | additionalFeatures, | ||
| const bool | useRetentionTimes, | ||
| std::vector< std::string > & | filenames, | ||
| std::vector< bool > & | exists | ||
| ) | [static] |
Returns a list of the Percolator input and output files for the specified data file (deprecated).
This static function can be called without creating an ms_mascotresfile object, and can be used in advance of creating an object to see if the percolator files will need to be created. If an object has already been created, it is normally easier to call setPercolatorFeatures() and then getPercolatorFileNames()
See Using Percolator scores and Using STL vector classes vectori, vectord and VectorString in Perl, Java, Python and C#.
The offsets into the arrays are defined by ms_mascotresfile::PERCOLATOR_FILE_NAMES
| szDatFileName | is the absolute or relative path to the results file |
| cacheDirectory | will normally be the value returned from ms_mascotoptions::getCacheDirectory |
| percolatorFeatures | is normally retrieved by calling ms_mascotoptions::getPercolatorFeatures(). The filenames encode the features so that there is no conflict |
| additionalFeatures | is normally a string passed to ms-createpip.exe For example, "-a numUniqPeps -r varmods" would add numUniqPeps and remove varmods from the default. Any other parameters except -a and -r are ignored. |
| useRetentionTimes | is a flag to indicate whether retention time information is used by percolator.exe |
| filenames | returns the list of files |
| exists | is a boolean vector which will return flags indicating if the Percolator files exists. The first value in the vector corresponds to the first value in the filename vector. |
| bool staticGetPercolatorFileNames | ( | const char * | szDatFileName, |
| const char * | cacheDirectory, | ||
| const ms_mascotoptions & | options, | ||
| const char * | additionalFeatures, | ||
| std::vector< std::string > & | filenames, | ||
| std::vector< bool > & | exists | ||
| ) | [static] |
Returns a list of the Percolator input and output files for the specified data file.
This static function can be called without creating an ms_mascotresfile object, and can be used in advance of creating an object to see if the percolator files will need to be created. If an object has already been created, it is normally easier to call setPercolatorFeatures() and then getPercolatorFileNames()
Make sure you set PercolatorExeFlags in options based on whether the results file has any queries with a retention time. Otherwise, this method may generate a filename different from setPercolatorFeatures().
bool anyRT = (whether any query has RTINSECONDS);
std::string percolatorFlags = options.getPercolatorRtFlags(anyRT, options.isPercolatorUseRT());
options.setPercolatorExeFlags(percolatorFlags.c_str());
See Using Percolator scores and Using STL vector classes vectori, vectord and VectorString in Perl, Java, Python and C#.
The offsets into the arrays are defined by ms_mascotresfile::PERCOLATOR_FILE_NAMES
| szDatFileName | is the absolute or relative path to the results file |
| cacheDirectory | will normally be the value returned from ms_mascotoptions::getCacheDirectory |
| options | contains the options stored in mascot.dat It is used to access the relevant options to generate the file names |
| additionalFeatures | is normally a string passed to ms-createpip.exe For example, "-a numUniqPeps -r varmods" would add numUniqPeps and remove varmods from the default. Any other parameters except -a and -r are ignored. |
| filenames | returns the list of files |
| exists | is a boolean vector which will return flags indicating if the Percolator files exists. The first value in the vector corresponds to the first value in the filename vector. |
| bool versionGreaterOrEqual | ( | int | major, |
| int | minor, | ||
| int | revision | ||
| ) | const |
Compare the value returned by getMascotVer() with the passed version number.
Utility function to perform easy comparison. For example, to test if a results file could have taxonomy information, use:
if (versionGreaterOrEqual(2, 4, 0)) then ...
| major | is the major version to be compared with |
| minor | is the minor version to be compared with |
| revision | is the minor revision to be compared with |
| bool willCreateCache | ( | const char * | szFileName, |
| const ms_mascotoptions & | opts, | ||
| const char * | applicationName, | ||
| std::string & | resfileCacheFileName, | ||
| unsigned int & | cacheStatus | ||
| ) | [static] |
Returns true if a cache file will be created when the ms_mascotresfile constructor is called.
This static function can be called without creating an ms_mascotresfile object. It can be used in advance of creating the object to see if there will be a delay while (re)creating the cache file(s).
The purpose of this method is to get the status of the cache in addition of whether the cache will be created or not.
See Multiple return values in Perl, Java, Python and C#
See Static functions in Perl, Java, Python and C#
| [in] | szFileName | is the absolute or relative path to the Fxxxxx.dat file |
| [in] | opts | normally loaded from the mascot.dat file using ms_datfile::getMascotOptions() |
| [in] | applicationName | is the name of the application or script that is calling this function. The applicationName is searched for in the return value from ms_mascotoptions::getResultsCache and ms_mascotoptions::getResfileCache to determine if the application should be using cache files. If it is not found then the function returns false and sets the cacheStatus to ms_peptidesummary::RESFILE_CACHE_DISABLED_IN_OPTIONS. If null, or an empty string is passed, no check is made. |
| [out] | resfileCacheFileName | returns the name of the ms_mascotresfile cache file if one exists or would be created |
| [out] | cacheStatus | is the ms_peptidesummary::CACHE_STATUS enumeration which gives more details about why the cache file may or may not be created. Multiple values may be bitwise OR'd toegether. |
| bool willCreateCache | ( | const char * | szFileName, |
| const unsigned int | flags, | ||
| const char * | cacheDirectory, | ||
| std::string * | cacheFileName | ||
| ) | [static] |
Returns true if a cache file will be created when the ms_mascotresfile constructor is called.
This static function can be called without creating an ms_mascotresfile object, and can be used in advance of creating an object to see if there will be a delay while (re)creating a cache file. The function has the same parameters as the ms_mascotresfile constructor - see the documentation for that function for details
See Static functions in Perl, Java, Python and C#
See Multiple return values in Perl, Java, Python and C#.
| [in] | szFileName | - see ms_mascotresfile::ms_mascotresfile |
| [in] | flags | - see ms_mascotresfile::ms_mascotresfile |
| [in] | cacheDirectory | - see ms_mascotresfile::ms_mascotresfile |
| [out] | cacheFileName | - the full path name of the cache file. For languages other than C++, this will be a reference rather than a pointer to a std::string. |
| Copyright © 2022 Matrix Science Ltd. All Rights Reserved. Generated on Thu Mar 31 2022 01:12:34 |