|
To view this email as a web page, click here. |
 |
|
Welcome
Thank you to all who visited us in Indianapolis at the annual ASMS conference.
If you weren't able to attend our breakfast workshops, you can
view the presentations here.
We are delighted to announce that Mascot Server 2.6.1 is now available.
The Mascot Server search result export utility is very powerful and we have some suggestions on how to get the most out of it.
This month's highlighted publication shows a new method for the detection of antibiotic-resistant strains of bacteria.
If you have a recent publication that you would like us to consider for an upcoming Newsletter, please
send us a PDF or a URL.
Mascot tip of the month explains how to automate a housekeeping task to conserve free disk space on your Mascot Server.
Please have a read and feel free to contact us if you have any comments or questions. |
|
|
|
 |
 |
 |
|
Mascot Server 2.6.1
Mascot 2.6.1 has been released and updates from 2.6.0 for both Windows and Linux can be
downloaded from the support page.
This patch release includes a number of important bug fixes, and we advise all users to install it. In particular, it fixes some spectral library issues: working with library files over 2 GB, using annotations to improve scores, and exporting of search results that include library matches. The procedure to create a library from existing search results is now more flexible and resilient.
|
 |
 |
 |
 |
|
Exporting search results
The Mascot Server export utility allows search results to be exported in a wide range of formats, together with the native result file and the MGF peak list.
This doesn't require file share access to your Mascot Server. The export utility can be used interactively, in a web browser, by following the appropriate link in a result report, or it can be executed automatically by Mascot Daemon, after each search in a task.
If you have command line access to your server, you can also export a set of result files by creating a Windows batch file or a Linux shell script. This latter approach is rather involved and reading more on this is recommended.
Useful hints concerning some less obvious aspects of exporting can be found in this posting.
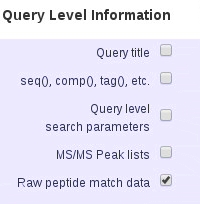
- To include modification site analysis in the CSV and XML export formats, check Query Level Information and Raw peptide match data. It is then output as a separate section, after the main protein and peptide table.
- If you want to export MS2 quantitation results (reporter or multiplex) you need to check the quantitation boxes under both Protein Hit Information and Peptide Hit Information.
- Export of spectral library search results is now available in version 2.6.1. If you are running 2.6.0, you can download a service pack from the support page.
|
 |
 |
 |
 |
|
Featured publication using Mascot
Here we highlight a recent interesting and important publication that employs Mascot for protein identification, quantitation, or characterization. If you would like one of your papers highlighted here please send us a PDF or a URL.
|
|
|
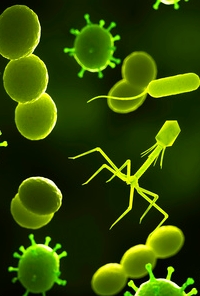
Detection of methicillin-resistant Staphylococcus aureus using phage amplification combined with matrix-assisted laser desorption/ionization mass spectrometry
Jon C. Rees, John R. Barr
Anal. Bioanal. Chem. (2017) 409: 1379
The US Centers for Disease Control and Prevention recently reported on the threats posed by antibiotic resistant bacteria. They estimated that each year in the USA, 2 million people become infected and at least 23,000 people die from resistant bacteria, at a healthcare cost of $20 billion. They also stated that this is a worldwide problem.
To enable faster detection of these resistant strains, the authors have developed a method where samples with S. aureus are incubated together with bacteriophage. They are then rapidly digested with trypsin and analyzed by MALDI-TOF MS. The method detects tryptic peptides from amplified phage capsid proteins, and has more sensitivity than methods that have tried to detect the intact phage proteins.
Antibiotic resistance is determined by the detection of phage replication in the presence of an antibiotic, when compared to a no-antibiotic control. After treating four S. aureus strains with cefoxitin, significant phage amplification in both the no-antibiotic and the cefoxitin-containing samples was shown on the resistant strains, while the susceptible strains showed no amplification upon cefoxin treatment.
|
 |
 |
 |
 |
|
Mascot tip of the month
It isn't uncommon for the disk on a Mascot Server to become full and for searches to then fail. Much of the space is taken up by the results (.dat) files and the space required can be reduced by compressing all old results files. All of the programs on the Mascot Server that need to read a results file will un-compress a file if necessary. So, if you want to go back and look at an old results file that has been compressed, it will just take a bit longer to open.
Mascot uses cache files to speed up report loading and navigation and by default these files are saved in the 'cache' subdirectory under the data directory. If you delete these files, then Mascot will re-create them when you look at a report, so it's totally safe to delete all of them.
You can obviously do both of these manually, but it makes more sense to automate this process. With Mascot Server 2.6, we included a script to delete old files in the cache directory and to compress the results (.dat) files. By default, this script isn't enabled, but it's very easy to enable it. If you open the mascot.dat file (in, for example the \inetpub\mascot\config directory) using Windows notepad or your favourite text editor, and scroll to the bottom, you will see a line like this:
# 0 0 * * 6 C:\inetpub\mascot\perl64\bin\perl.exe C:/inetpub/mascot/bin/tidy_data.pl --clearcache --compressdat
The paths may be different on your system and you shouldn't need to change them. To enable the script, you need to remove the # and the space from the start of the line so the first character on the line is the '0'. The first 6 values specify when the script should be run. The first '0' means 0 minutes past the hour, the second '0' means midnight. The '*' and '*' means any day of the month and any month of the year. The '6' means the sixth day of the week, which is defined as Saturday. By default, the script will only process files created more than 100 days ago, but you can change this by adding ‑‑minage=100 to the end of the line and replacing the 100 with a different value. When the script is run, it writes to a file, tidy_data.log in the mascot logs directory.
If you wish to use this script with Mascot 2.4 or Mascot 2.5, see page 33 of
this presentation for details.
|
 |
 |
 |
 |
|
About Matrix Science
Matrix Science is a provider of bioinformatics tools to proteomics researchers and scientists, enabling the rapid, confident identification and quantitation of proteins. Mascot software products fully support data from mass spectrometry instruments made by Agilent, Bruker, Sciex, Shimadzu, Thermo Scientific, and Waters.
Please contact us or one of our marketing partners for more information on how you can power your proteomics with Mascot.
|
 |
 |
|
|