|
To view this email as a web page, click here. |
|
|
|
Welcome
Mascot Server can now search spectral libraries in BiblioSpec BLIB format by using the new conversion script blib2msp.
This month's highlighted publication examines PET plastic degradation using a PET hydrolase expressed in fruit flies.
Our London office has moved! Please update your address book.
|
|
|
|
|
|
 |
|
Mascot: The trusted reference standard for protein identification by mass spectrometry for 25 years
|
Get a quote
|
 |
|
|
|
Converting BLIB spectral libraries to NIST MSP
|
|
|
Mascot Server can create and search NIST MSP formatted spectral libraries.
However, many spectral libraries use a different file format and they are not directly compatible.
One example is BiblioSpec, which is part of the Skyline software and uses the BLIB format.
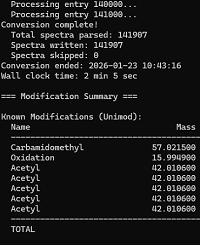
We have developed a script, blib2msp, for converting BLIB to MSP files.
This means Mascot Server can now search DDA libraries generated by Skyline (BiblioSpec) as well as tools that generate in silico libraries like Carafe.
You can also use blib2msp to convert NIST MSP formatted files generated by Mascot Server into BLIB.
Skyline can already read NIST (.MSP) files, but MSP libraries converted to BLIB format may be useful to other spectral library search engines.
It will allow one library to be used consistently across multiple platforms whatever the source.
A full description as well as link to the Matrix Science GitHub repository is in our blog.
|
|
|
|
 |
|
|
|
Featured publication using Mascot
Here we highlight a recent interesting and important publication that employs Mascot for protein identification, quantitation, or characterization. If you would like one of your papers highlighted here, please send us a PDF or a URL.
|
|
|
Polyethylene terephthalate degradation by Drosophila melanogaster through heterologous expression of glycosylated polyethylene terephthalate hydrolase (PETase)
Rikako Sanuki, Hatsune Minami, Emi Kawano, Keita Katsuma, Toshiyuki Takano-Shimizu-Kouno, Shosuke Yoshida
Communications Sustainability volume 1, Article number: 36 (2026), doi:10.1038/s44458-026-00047-5
Recycling the widely used plastic Polyethylene terephthalate (PET) typically relies on energy-intensive chemical processes.
PET hydrolytic enzymes, such as PET hydrolase (PETase) from Piscinibacter sakaiensis, have been proposed as an alternative allowing for biological PET recycling.
The authors explored using insects as a platform for PET biodegradation by genetically engineering Drosophila melanogaster to express PETase in a subset of the intestinal tract and salivary glands, demonstrating degradation of both a water-soluble PET copolymer and of solid PET.
PETase expressed in both D. melanogaster and human cells was found to be glycosylated, which reduces catalytic activity but increases enzyme stability, allowing sustained PET hydrolysis over longer periods.
To profile N-glycosylation, the authors performed LC–MS/MS on tryptic and chymotryptic peptides following deglycosylation. The resulting spectra were searched using Mascot Server to detect deglycosylation-induced mass shifts caused by conversion of glycosylated asparagine residues to aspartate following enzymatic treatment. Deamidated Asn residues within the Asn-X-Ser/Thr motif were considered N-glycosylation sites. This enabled confident identification of N-glycosylation sites within the PETase sequence and confirmed that specific residues were modified in the eukaryotic expression systems.
|

|
|
|
 |
|
|
|
Change of address for UK office
|
|
|
Matrix Science Ltd is based in London, UK since 1998. Until recently, our UK office was at 64 Baker Street.
We have moved to new premises one block up the street. The new address is:
Matrix Science Ltd
83 Baker Street
London, W1U 6AG
United Kingdom
There is no other change to our contact details, and our phone numbers and email addresses remain the same.
|

|
|
|
 |
|
|
|
About Matrix Science
Matrix Science is a provider of bioinformatics tools to proteomics researchers and scientists, enabling the rapid, confident identification and quantitation of proteins. Mascot continues to be cited by over 2000 publications every year. Our software products fully support data from mass spectrometry instruments made by Agilent, Bruker, Sciex, Shimadzu, Thermo Scientific, and Waters.
Get a quote
|
|
|
You can also contact us or one of our marketing partners for more information on how you can power your proteomics with Mascot.
|




|
|
|
|
|