|
To view this email as a web page, click here. |
|
|
|
Welcome
Mascot Server can create and search DDA spectral libraries. We illustrate this with a custom Mycobacteria library.
This month's highlighted publication establishes orthogonal translation systems for lysine aminoacylation.
We are presenting three posters about Mascot DIA at ASMS 2026 in June.
|
|
|
|
|
|
 |
|
Mascot: The trusted reference standard for protein identification by mass spectrometry for 25 years
|
Get a quote
|
 |
|
|
|
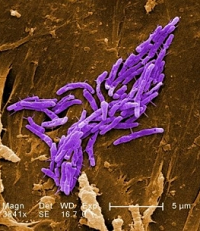
Identifying Mycobacteria with custom spectral libraries
|
|
|
Mass spectrometry can be used for rapid identification of difficult-to-culture or slow-growing bacteria so that a patient can receive prompt treatment.
The quickest identification method is searching the MS/MS spectra against a species-specific spectral library (SL), which is supported in Mascot Server.
Chinmaya Kotimoole et al. (2024) analysed proteomes of seven nontuberculous mycobacteria, gathered from 26 data sets and totaling more than 20 million peptide spectrum matches.
A variety of instruments and search engines were used to analyse or reanalyse the DDA data, and spectral libraries were created in Skyline in the BLIB format.
We selected a number of them and used our blib2msp converter to transform them into the MSP format, required by Mascot Server.
The spectral libraries were not 'pure' as they contained a mixture of Mycobacteria, a mixture of human proteins, BSA and associated bovine proteins from the culture media.
We used blib2msp to extract all the peptide sequences from the SL, then processed them as an assay in Unipept, which maps peptides to species.
We filtered each list to a single Mycobacteria species and used this as a filter with blib2msp to create species-specific libraries.
To demonstrate, we used data from a different Mycobacteria project (PRIDE project PXD059923) and searched them against the filtered spectral libraries.
The most useful analysis mode is to search a FASTA sequence database plus a species-specific SL, which is implemented by Mascot Server as the integrated library search.
We were able detect several species by their unique peptides in the test data set.
The full steps and results are in our blog.
|
|
|
|
 |
|
|
|
Featured publication using Mascot
Here we highlight a recent interesting and important publication that employs Mascot for protein identification, quantitation, or characterization. If you would like one of your papers highlighted here, please send us a PDF or a URL.
|
|
|
Expanding the Genetic Code with Lysine Aminoacylation
Xinyu Li, Qinglei Gan, Chenguang Fan
Journal of the American Chemical Society, 2026, 148, 11, 12411–12422, doi:10.1021/jacs.6c03157
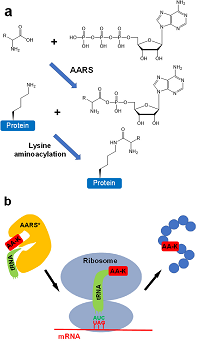
Lysine aminoacylation is a recently discovered post-translational modification in humans, which remains poorly studied due to the difficulty of generating homogeneously aminoacylated proteins at target lysine sites.
The authors establish orthogonal translation systems (OTS) for ten different types of lysine aminoacylation, compatible with bacterial and mammalian cells.
They use a genetic code expansion strategy, where aminoacyl-tRNA synthetase (AARS) is engineered to recognize an aminoacylated lysine (AA-K), and an engineered tRNA – usually with a mutated anticodon – decodes an assigned codon.
The authors use the method to demonstrate the effect of lysine aminoacylation on essential metabolic enzymes and cellular metabolism both in vitro and in vivo.
AA-K-containing superfolder green fluorescent protein (sfGFP), AA-K-containing pyruvate kinase M2 (PKM2) or glucose-6-phosphate dehydrogenase (G6PD), acetyllysine-containing PKM2 or G6PD and Val-K-containing PKM2 were expressed and purified.
The samples were separated by SDS-PAGE and analysed with LC-MS/MS (Thermo LTQ Orbitrap XL), then the DDA spectra searched with Mascot Server to identify and localise the modified lysines.
The structure of PKM2 variants was additionally analysed with circular dichroism (CD) spectrometry, and the glyolytic rate measured with a Seahorse XF glycolytic rate assay.
|
|
|
|
 |
|
|
|
Matrix Science at ASMS 2026
|
|
|
Matrix Science is attending ASMS 2026 this June in San Diego, CA.
Please drop by booth #616.
We have been working hard on finalising DIA support in Mascot.
At ASMS 2026, we are presenting three posters featuring Mascot DIA using real data.
Mascot DIA introduces probabilistic scoring to spectrum-centric analysis of DIA data.
Presented by Ville Koskinen.
Thursday 3 June, Informatics: Peptide ID and Quantification.
Mascot DIA enables direct comparison of DDA and DIA identification and quantitation results.
Presented by Patrick Emery.
Thursday 3 June, Informatics: Peptide ID and Quantification.
Overcoming Peptide-Centric DIA Limitations: Discovering Unsuspected Modifications in Narrow-Window DIA Using Mascot Error Tolerant Search.
Presented by Richard Jacob.
Thursday 3 June, Peptides: PTM Identification
|

|
|
|
 |
|
|
|
About Matrix Science
Matrix Science is a provider of bioinformatics tools to proteomics researchers and scientists, enabling the rapid, confident identification and quantitation of proteins. Mascot continues to be cited by over 2000 publications every year. Our software products fully support data from mass spectrometry instruments made by Agilent, Bruker, Sciex, Shimadzu, Thermo Scientific, and Waters.
Get a quote
|
|
|
You can also contact us or one of our marketing partners for more information on how you can power your proteomics with Mascot.
|




|
|
|
|
|