Sequence database setup: IPI
|
Overview
NOTE: IPI is now obsolete. EBI announced they would cease maintaining the IPI databases in 2011. The suggested alternative is UniProt Proteomes.
IPI (International Protein Index) is compiled by the EBI (European Bioinformatics Institute) to provide a top level guide to the main databases that describe the proteomes of the higher eukaryotic organisms. The aim is to:
- effectively maintain a database of cross references between the primary data sources
- provide a minimally redundant yet maximally complete set of proteins (one sequence per transcript)
- maintain stable identifiers (with incremental versioning) to allow the tracking of sequences in IPI between IPI releases.
There are seven IPI databases, Homo sapiens, Mus musculus, Rattus norvegicus, Danio rerio, Arabidopsis thaliana, Gallus gallus, and Bos taurus. This document uses the Human database as an example. To work with the other database, simply substitute the name of the organism. For example, the compressed Fasta file for Mus musculus is ipi.MOUSE.fasta.gz, the db_update.pl keyword is IPI_mouse_from_EBI, the recommended Mascot name is IPI_mouse, etc.
Download
ftp://ftp.ebi.ac.uk/pub/databases/IPI/current/
for the latest release.
ftp://ftp.ebi.ac.uk/pub/databases/IPI/old/
for earlier releases.
There are two files: a Fasta database file (ipi.HUMAN.fasta.gz) and a reference file in Swiss-Prot format (ipi.HUMAN.dat.gz). It is worth getting the reference file because then you can view a full text report, including cross reference information, without linking out to the internet.
Taxonomy
Taxonomy is not required because all entries are from the same species
Parse Rules
A typical Fasta title line is:
>IPI:IPI00177321.1|SWISS-PROT:Q5JTD7|TREMBL:B3KX61;Q3B825|ENSEMBL:ENSP00000361518|REFSEQ:NP_001012992|H-INV:HIT000339065|VEGA:OTTHUMP00000016460 Tax_Id=9606 Gene_Symbol=C6orf154 Uncharacterized protein C6orf154
The IPI accession number is the preferred identifier. In most cases, it is not necessary to include the version number.
Accession from Fasta title: ">IPI:\([^| .]*\)"
Description from Fasta title: ">[^ ]* \(.*\)"
The corresponding line in the Dat file is:
ID IPI00177321.1 IPI; PRT; 316 AA.
Accession from Ref file: "^ID \([^ .]*\)"
Configuration (Mascot 2.3 and earlier)
For this example, both database files were downloaded to C:\Inetpub\MASCOT\sequence\IPI_human\current, decompressed using gzip, and renamed to IPI_human_3.61.dat and IPI_human_3.61.fasta.
When updating an active database, it is important to rename the Fasta file last, because Mascot will begin database exchange as soon as it sees a new Fasta file that matches the wildcard path for the database.
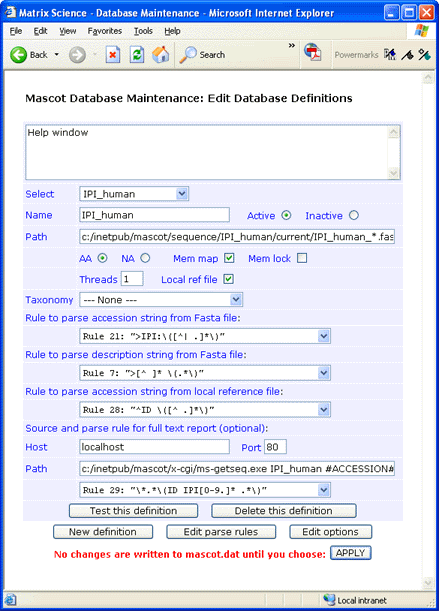
If you don’t require full text in a Mascot Protein View report, simply leave the Host, Port, and Path fields blank
and choose
— no full text report —
in the drop down list.
Always test a new definition before applying the changes to mascot.dat