Peak picking intact crosslink spectra with Mascot Distiller
There are several important factors to take into account when carrying out peakpicking for an intact crosslinked dataset. You’ll typically be dealing with higher charge state precursors to handle the larger masses of the linked peptides, and the MS/MS spectra are inherently complex, chimeric spectra with fragments from the alpha and beta peptides. To look at the effects of adjusting peak-picking settings we took the same raw file from the publicly available synthetic peptide library crosslink dataset used in the Validating intact crosslinked peptide matches blog article and processed with Mascot Distiller 2.8.3 using a range of different processing settings.
Fragment ion charge states
Mascot Server only searches for 1+ and 2+ fragment ion series but if you have higher charge state precursors, as you are likely to have with a crosslinked dataset, many of the fragment ions generated will similarly have higher charge states and so can’t be matched (or worse, might result in false positive matches). With these types of datasets, you should either de-charge the fragment ion peaks to MH+, or, if you have Mascot 2.7 or later installed, supply the peak charge states. Mascot Distiller can do this, but if your MS/MS scans are saved as centroids, rather than profile, then you need to use the uncentroiding options.
To examine this, we carried out peak picking with Mascot Distiller 2.8.3 using the following processing settings:
- default.ThermoXcalibur.opt, which simply take the centroid values from the raw file without determining charge state
- the settings used in the Validating intact crosslinked peptide matches blog article, which uncentroid the MS/MS scans at a resolution of 100 points per Dalton (ppDa)
- prof_prof.ThermoXcalbur.opt, which uncentroid the MS/MS scans at a resolution of 600 ppDa
| Processing options | Uncentroiding points per Da | Decharged fragment ions? | Time taken (mm:ss) | Significant CSMs (p<0.05) |
|---|---|---|---|---|
| default.ThermoXcalibur.opt | N/A | No | 00:14 | 655 |
| Previous blog settings | 100 | Yes | 02:34 | 1087 |
| prof_prof.ThermoXcalibur.opt | 600 | Yes | 12:08 | 1143 |
One thing to notice is that if you compare the results using the processing options from the previous blog article, we are seeing more significant cross-linked spectrum matches (CSMs) – 1087 vs 1008 in the original article. The original article used Mascot 2.7 to search the data, and we have made some improvements to the crosslinking search in Mascot 2.8. These improvements have yielded an additional 79 matches.
More generally, the results from default.ThermoXcalibur.opt are significantly worse than with uncentroiding enabled. The reason for this is because we have many peaklists containing fragment ions with charge states in excess of 2+, which will not be matched by Mascot unless we decharge the fragment ions in the peaklists. For example, Figure 1 below shows the same cross-link match from a 4+ precursor using the default.ThermoXcalibur.opt and prof_prof.ThermoXcalibur.opt peak processing settings:
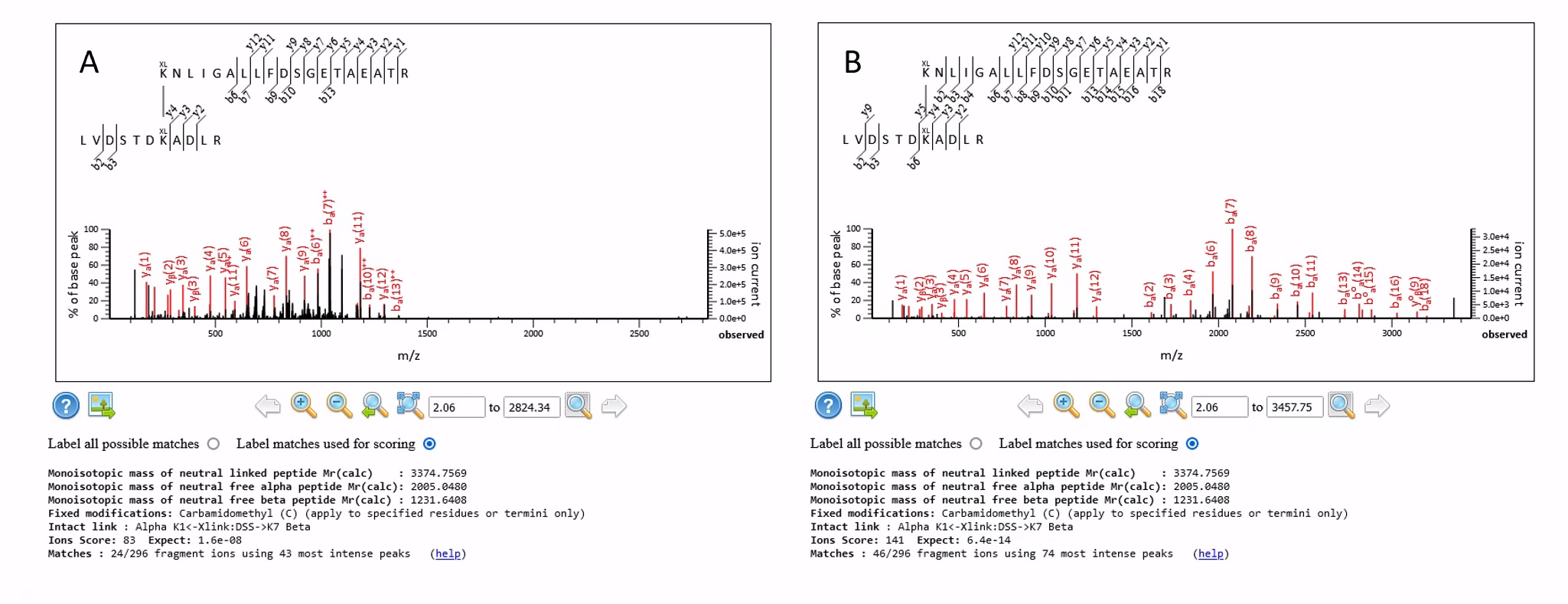 Figure 1 The same match where the data were processed using A) default.ThermoXcalibur.opt and B) prof_prof.ThermoXcalibur.opt and then decharged
Figure 1 The same match where the data were processed using A) default.ThermoXcalibur.opt and B) prof_prof.ThermoXcalibur.opt and then decharged
Both are strong matches, but we can see from the Mascot Distiller peaklist for the spectrum that there are a reasonable number of 3+ fragment ions (and a few 4+), so the decharged peak list from the prof_prof.ThermoXcalibur.opt settings is giving us increased coverage of both the alpha and beta peptides because those higher charge state fragments can now be matched, increasing the coverage of the fragment ions from 24 to 46 and improving the score from 83 to 141.
Uncentroiding resolution
While the prof_prof.ThermoXcalibur.opt settings out-performed the settings used in the previous blog article, finding an additional 56 significant CSMs, this comes at a heavy cost in processing time – taking approximately 5 times as long to complete. Previously we demonstrated that decreasing the uncentroiding setting in the prof_prof option to 200 or 400 ppDa could still give good results while improving processing time, so we also tried this on the crosslink data. Results are summaries in table 2 below:
| Uncentroiding ppDa | Time taken (mm:ss) | Significant CSMs (p<0.05) |
|---|---|---|
| 200 | 02:33 | 1139 |
| 400 | 06:25 | 1141 |
| 600 | 12:08 | 1143 |
As you can see, using either 200 or 400 ppDa both yield results close to the default of 600 ppDa, while taking approximately 1/5th and ½ the processing time. The reasons for these differences are the same – uncentroiding to a higher resolution improves peak picking and reduces the risk of nearby adjacent centroids being merged into a single uncentroided peak, so we find a few additional marginal CSMs compared with uncentroiding to 200 or 400 ppDa, but doing this requires more processing time. Reducing the uncentroiding resolution to 400ppDa seems to be a reasonable trade off between speed of processing and the quality of the results. Of course, if your data are saved as profile, rather than centroids, then this is not a trade-off you’ll need to make.
Signal to noise
Good signal to noise in the peaklist will generally result in a higher confidence match back from the database search. MS/MS spectra from intact crosslinked peptides are inherently complex chimeric spectra, making the signal to noise of the peaklists even more important. Figure 1 above shows that carrying out peakpicking in Mascot Distiller instead of simply taking the centroids is giving us cleaner peaklists with better signal to noise. However, it’s also clear that the default signal to noise cut off of 1 may be too low for these data, so we reprocessed the peaklists using 400 ppDa for uncentroiding with an increased value of 2 for the “Minimum signal to noise (S/N)” setting on the MS/MS Peak Picking options tab in Mascot Distiller. Results are summarised in table 3:
| Uncentroiding ppDa | Minimum signal to noise | Time taken (mm:ss) | Significant CSMs (p<0.05) |
|---|---|---|---|
| 400 | 1 | 06:25 | 1141 |
| 400 | 2 | 05:24 | 1152 |
For these data, increasing the signal to noise threshold has yielded an additional 11 CSMs, and reduced the processing time by 1/6th. What threshold to use on your own data will depend on the data produced by your instrument – setting too high a threshold will result in correct fragment ions being excluded from the peaklists, resulting in worse database search results. Our advice is to run a standard sample through your workflow, adjust the processing options to give them best combination of results and processing time for your data, and then use those as your default settings for processing with Distiller.
We’ve used data from a Thermo instrument in this blog, where MS/MS scans are routinely saved as centroids. Many non-Thermo instruments default to collecting data in profile mode, so you won’t need to uncentroid, but otherwise the points about decharging the fragment ions and adjusting the signal to noise thresholds apply.
If you don’t have a copy of Mascot Distiller and you’d like to evaluate it on your own data, we offer a 30 day trial of Mascot Distiller. For details, please see the Mascot Distiller download page.
Keywords: crosslinking, Mascot Distiller, peak picking, Thermo