How many of you are there in there? Processing and searching chimeric MS/MS spectra with Mascot Distiller and Mascot Server
In the typical shotgun proteomics experiment, the assumption is that each MS/MS spectrum is derived from a single precursor selected by the Mass Spectrometer for fragmentation. However, in practice, near isobaric precursors can co-elute and undergo co-fragmentation resulting in chimeric MS/MS spectra containing fragments from multiple different precursor peptides.
With high resolution data, it is possible for these overlapping isotope distributions to be cleanly resolved, as shown in Figure 1 below:
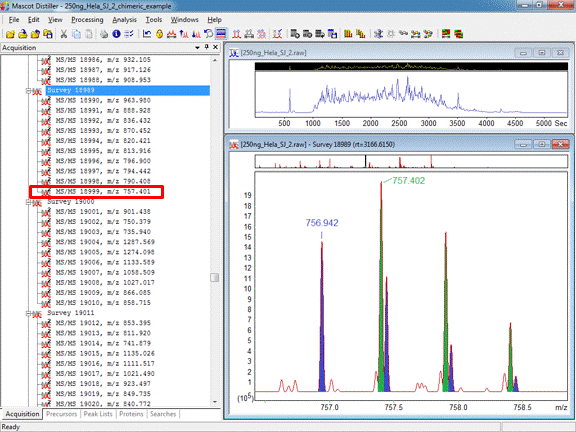 Figure 1: Overlapping isotope distributions
Figure 1: Overlapping isotope distributions
In Mascot Distiller 2.5, peak picking was extended to return multiple precursor m/z values in such cases, up to a user specified limit. The identified precursor masses can then be output as multiple PEPMASS parameters for each MS/MS peak list the exported MGF peak list file. To enable this, you need to ensure that following options are set prior to carrying out peak detection and generation of the peak lists file:
- 1. On the MS/MS Processing tab of the ‘Processing Options’ dialog, ensure that the ‘Maximum number of precursor m/z values’ is set to a value greater than 1.
- 2. Ensure that the ‘Allow multiple precursors per scan’ checkbox is checked on the ‘Peak List Format’ tab of the general preferences dialog.
In Mascot Server 2.5 and later, these sets of m/z values are treated collectively, with the MS/MS data being searched using each precursor mass value. If two precursors return different matches, both are reported using different query numbers. If two precursors return the same match, this is only reported once, using the closest m/z value.
To verify this multiple precursor approach, we took a publicly available yeast dataset from the PeptideAtlas repository, [PASS00665]. This dataset was used by Shteynberg et al. to validate reSpect [Shteynberg et al., JASMS 26(11): 1837-1847], a tool which can be used to identify additional matches from chimeric spectra. We carried out peak detection using Mascot Distiller, allowing for up to 4 precursors per MS/MS spectrum. The peak lists were then exported in the MGF format with and without multiple precursor masses enabled and searched using Mascot 2.5 against the S.cerevisiae sequences in the SwissProt database. Decoy searches were automatically preformed using the integrated decoy option in Mascot, and the significance thresholds adjusted to give 1% peptide FDR.
Figures 2 and 3 below show two peptide matches to the same MS/MS spectrum taken from the multiple precursor search. The matches are both significant and were found using two different precursor masses and charge states:
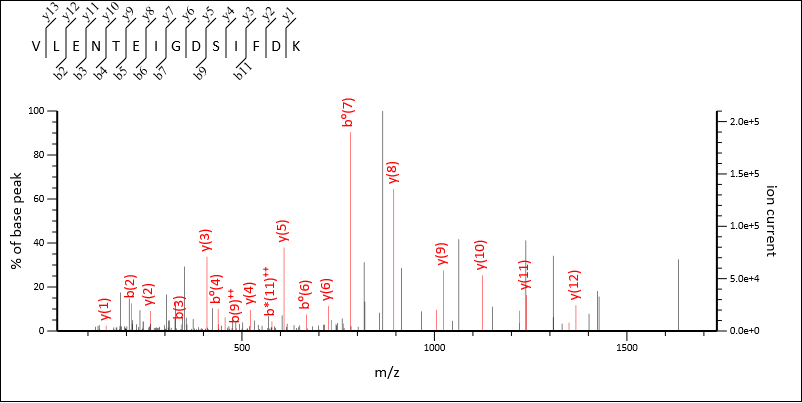 Figure 2: Match to MS/MS spectrum using a 2+ precursor with an m/z ratio of 790.396
Figure 2: Match to MS/MS spectrum using a 2+ precursor with an m/z ratio of 790.396
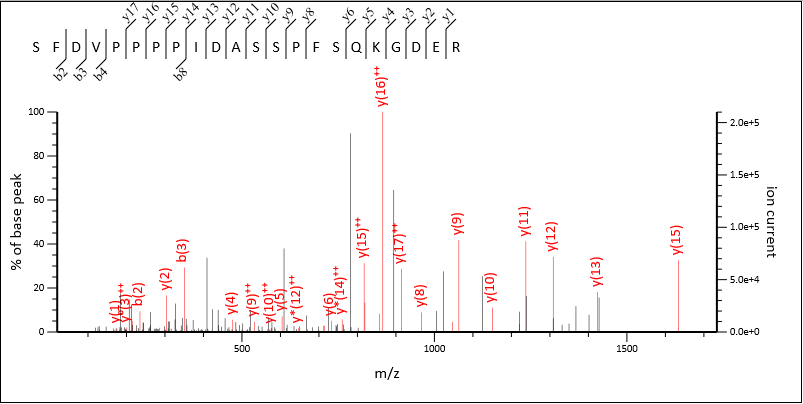 Figure 3: Match to MS/MS spectrum using a 3+ precursor with an m/z of 791.715
Figure 3: Match to MS/MS spectrum using a 3+ precursor with an m/z of 791.715
As you can see, there is a clear separation of the majority of the MS/MS fragments used for each match, and between them the majority of the most intense peaks in the spectrum have been assigned.
Table 1 summarises the statistics for the multiple precursor dataset as a whole. There were 342827 MS/MS spectra in total. On average, Mascot Distiller found just under 2 possible precursors for each spectrum, giving approximately double the number of search queries. The significance threshold was adjusted to give a 1% peptide false discovery rate, giving us 207882 queries with significant peptide matches identified, compared with 167806 from the single precursor search. Of these matches, 19117 were additional matches from chimeric spectra where we have two or more significant matches to the same MS/MS spectrum but from different precursors. Perhaps even more significantly, there were 20959 cases where the most intense precursor from the MS spectrum failed to get a match, but a less intense precursor gave a significant match. Of course, some of these matches could have been found in the search without multiple precursors if we had used a wider precursor mass tolerance. However, this would have decreased the specificity of the search, resulting in an increased search time and significance thresholds.
| Number of MS/MS spectra | 342827 |
| Number of search queries | 672146 (e.g. average 2 precursors per spectrum) |
| Number of significant matches (1% FDR) | 207882 |
| Number of significant ‘chimeric’ matches | 19117 |
| Number of significant matches to less intense precursor | 20959 |
| Number of significant matches from single precursor search (1% FDR) | 167806 |
As you can see from these results, we identified significantly more peptide matches from the multiple precursor search, and that this dataset contains a relatively large number of chimeric spectra. Of course, this dataset was chosen to demonstrate the presence of chimeric spectra, and in general you wouldn’t expect to see such a large number of chimeric spectra. A more typical dataset can be see in this presentation from our 2014 ASMS User group meeting, where the rate of chimeric spectra is closer to 2%. In addition to the ‘true’ chimeric spectra matches, using the multiple precursor information identified by Distiller, we were able to get significant matches to additional MS/MS spectra without having to widen the precursor mass tolerance to account for a second, less intense, precursor. Therefore, using the options available in Mascot Distiller and Mascot Server 2.5 or later has therefore significantly improved our coverage of this dataset.
Keywords: chimeric spectra, Mascot Distiller, peak picking
Very nice feature !
Congrats.
Great to see that it is now implemented in Mascot too!