Results round-up for the ‘dark matter’ challenge
In June, we tried to harness the power of crowd-sourcing to explain some of the unidentified modifications found in open database searches. We selected 20 abundant and unassigned mass deltas from Supplementary Table 3 of the recent MSFragger paper from Alexey Nesvizhskii’s group at U. Michigan and offered prizes for the first credible explanations.
There were 35 unannotated deltas in the top 500 detected mass shift features listed in Supplementary Table 3. 11 of these were large negative values, which could only be artefacts or losses of amino acids. Two of the positive values had fairly likely explanations: 52.9128 could be Cation:Fe[III] (52.9115) and 151.9956 could be DTT adduct to cysteine (151.9966). We arbitrarily dropped two others, 3.0076 and 252.9798, to achieve a round number of 20 unknowns for the challenge.
The first successful identification came from Zoltan Szabo, who spotted that 23.958 was probably an aluminium cation (23.9581). Soon after, several other deltas were found to correspond to combinations of amino acid residues, usually including K or R. With the benefit of hindsight, the explanation was obvious: the MSFragger searches only allowed for one missed cleavage. If the matched peptide included two missed cleavages, this would be matched as a shorter peptide plus a modification. For example, K.SKLPKPVQDLIK.M would give a match as K.LPKPVQDLIK.M with an N-term mod of SK.
Martin Pabst suggested 306.0952 might be glycation (Addition of 2 Hexose and loss of water = C(12)H(18)O(9) = 306.0951). The only difficulty with this is the absence or near absence of similar modifications, such as addition of 1 hexose with loss of water or addition of 2 Hexose without loss of water. Bruce Onisko suggested the same composition but a different chemistry: modification by glucose then esterification with 3-hydroxy-3-methylglutarate. But, as with glycation, for any artefactual double modification, you would expect to see the individual modifications at higher abundance. In the HeLa dataset, the abundance of Hex is 78 PSMs / 22 peptides while the abundance of 306.0952 is 2812 PSMs / 766 peptides. In purely statistical terms, while C(12)H(18)O(9) may well be correct, it seems unlikely that 306 can be a secondary modification of Hex.
Similar considerations apply to 234.0742. Martin Pabst suggested Hex+carboxyethyl (C(9)H(14)O(7) = 234.0740) but carboxyethyl alone is not present in the top 500 features. Bruce Onisko proposed an alternative chemistry of modification by glucose then esterification with lactic acid, but the same reservations apply.
Bruce also commented offline that he believed 176.7462 contained no carbon. One possibility is Fe(3)H(-7)O = 176.7450, because we know that iron is very abundant, assuming 52.9115 is indeed Cation:Fe[III]. Alternatively, we could just say that 176.7462 looks like 3 x 52.9115 + water.
Some transition metal cations create special problems because of their isotope distributions. Peak picking tries to identify the monisotopic peak on the basis of the average elemental composition for peptides, which means the monisotopic peak is the first peak in the distribution. If a peptide is unlucky enough to pick up a mercury cation, the 2+ isotopic envelope for an 1800 Da peptide would look like the picture below. Cadmium would have a similar effect. Not much chance of assigning the monoisotopic mass correctly. So, when considering a composition that includes a transition metal, it may be necessary to allow for systematic mass errors of 1, 2 or 3 Da.
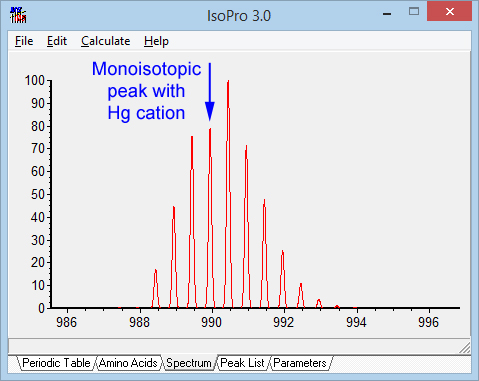
The other deltas remain unexplained. In particular, the two highly abundant but unidentified mass deltas originally reported in Steven Gygi’s 2015 paper: 301.9864 and 249.9803. This competition has certainly demonstrated that figuring out the chemistry for an unknown delta is very difficult.
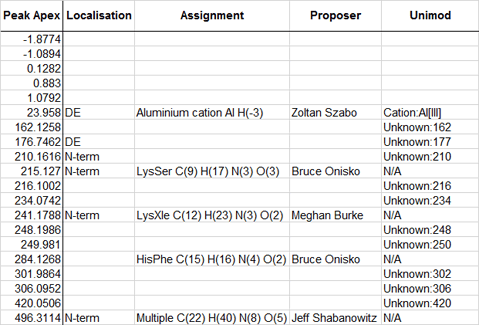
The positive deltas greater than 5 Da that were not in Unimod and were not due to missed cleavages have been added to Unimod with the prefix Unknown, e.g. Unknown:302. The elemental compositions of these entries are fakes, chosen to simulate the observed delta. The site specificities are also unknown, so they have been added with specificities of D, E, N-term, and C-term.
If you perform an error tolerant search and these deltas are present, strong matches will help narrow down the site specificity. It will be interesting to get feedback on site specificity and how often these modifications are observed because an error tolerant search is much more stringent than an open search, where only unmodified fragments are matched and there is no protection against reporting a spurious modification due to a mis-assigned precursor mass. Please add your feedback as comments to this blog article.
—
Note on calculating mass values with negative element counts from Bruce Onisko
Here is a method to find elemental formulas where elements are added and removed to achieve a specific high resolution mass modification.
To find such combinations of atoms, I have added the mass of C11H12N4O3S (which is 280.0630) to the mass of the unknown modification, searched for formulas that match to specific tolerances, then subtract C11H12N4O3S from the result. I use ChemCalc Molecular Formula finder with element ranges C0-50 H0-100 N0-4 O0-16 S0-4 P0-4, require unsaturations to be integers, and solutions to fit +/- 0.001 amu, since the data quality is excellent. Why C11H12N4O3S? This is a formula that allows the loss of any of the 20 common amino acids.
For example, if you try this for the known mass mod of 0.98402, the result is H-1N-1O, or –NH+O, which is not very recognizable. However an equivalent expression, –NH2+OH immediately suggests deamidation.
I used this method on the remaining unknowns in your table (and in Alexey’s), and here are some possible solutions, all better than 0.001 amu for three from the table and one more from Alexey’s Supplemental Table 3. (I ignore the other unknowns where more than 1 solution better than 0.001 amu was found.)
| Mod | C | H | N | O | P | S | or easier to see | loss of 1 AA? |
|---|---|---|---|---|---|---|---|---|
| 0.128 | 1 | 17 | -2 | 0 | 1 | -1 | -N2S +CH17P | ? |
| 1.079 | -1 | 13 | -1 | -1 | 2 | -1 | -CNOS+H13P2 | ? |
| -1.877 | 2 | 18 | -2 | -3 | 0 | 1 | -N2O3+C2H18S | -TrpCO2 + C14H28S |
| 3.008 | 2 | 5 | -3 | -1 | 0 | 1 | -N3O+C2H5S | -His + C8H14OS |
The 1st two solutions look impossible, the 3rd and 4th are reasonable, but no biochemistry comes to mind to explain them.
Keywords: ChemCalc, dark matter, delta mass, error tolerant, mass tolerant, modification, open search, Unimod
Judit Villen of U. Washington has figured out the likely explanation for the delta of 420.0506, which was present in TNBC at a very high level. It is probably an adduct of DTT+CAM dimer, C(12)H(24)N(2)O(6)S(4), that is lost quantitatively on MS/MS. When the survey scans are examined, there is perfect co-elution between the unmodified and modified peptide pairs. The calculated mass for the adduct is 420.05172 and the isotopic envelope for the protonated molecule, which can also be observed, is exactly as predicted. If you go looking for them, there is also evidence for adducts of CAM-DTT-CAM (268.05515) and CAM-DTT-DTT-DTT-CAM (572.04829), but at much lower abundance.