Thermo Scientific instruments
All Thermo Scientific instruments for proteomics use Thermo Xcalibur™ acquisition software. There is a wide choice of software to convert Xcalibur data into peak lists and submit these to a Mascot Server for searching. This page lists some of the more widely used options.
Note that spectra in Xcalibur raw files may contain centroid data, rather than profile data. With hybrid instruments, it is common for the high resolution survey scans to be profile and lower resolution MS/MS scans to be centroid.
Mascot Distiller
Mascot Distiller can be used to browse Xcalibur raw files, and process them into high quality peak lists that can be saved or submitted direct to a Mascot Server for searching. With the appropriate Distiller Toolboxes, the search results can be imported back into Distiller for further examination or used as the basis for quantitation. If the optional Mascot Daemon Toolbox is installed, these processes can be automated using Mascot Daemon.
If your MS/MS data is centroided, you can choose to create a peak list direct from the centroid values already present in the raw file. This is extremely fast and the peak list is fine for most purposes. Choose default.ThermoXcalibur.opt as the processing options when opening the raw file as a new project.
With high resolution data from an FT or Orbitrap, you may wish to take a little longer, and peak pick the survey scans, so as to obtain more reliable detection of the 12C peaks. For high charge state data, you may wish to peak pick the MS/MS scans so that the peaks can be de-isotoped and de-charged. Mascot only tries to match 1+ and 2+ fragments, so de-charging to 1+ becomes important when the precursor is 4+ or higher.
Please refer to Peak picking Thermo .RAW data with Mascot Distiller for full details.
Besides the quality of the peak lists, the other advantages of using Mascot Distiller are that it provides a universal interface to other raw file formats, and it is fully integrated with Mascot Server and Mascot Daemon. You can, for example, use Mascot Daemon to process batches of files automatically, saving Distiller project files that contain both the peak lists and the search results.
Mascot Daemon
Mascot Daemon can be used to process batches of RAW files by choosing Mascot Distiller as the data import filter.
Mascot Distiller requires the optional Mascot Daemon Toolbox to allow the Distiller libraries to be called from Mascot Daemon. When this toolbox is active, Mascot Distiller will appear automatically on the list of data import filters in Daemon.
See also Automated processing with Mascot Daemon real-time monitor.
Proteome Discoverer
Thermo’s Proteome Discoverer™ provides fully automated raw file processing and search submission to Mascot Server. Peak picking and search parameters are selected in a workflow wizard. When the search is complete, the results are imported into Proteome Discoverer, where they can be filtered and inspected.
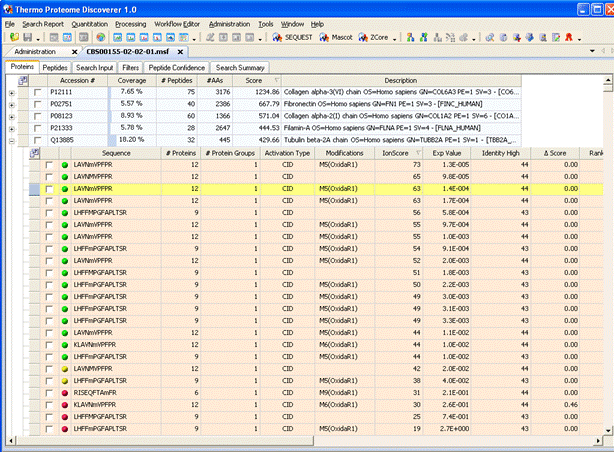
Note that the local Mascot Server URL must be entered in the form http://ec-vm2/mascot/ where ec-vm2 is replaced by the hostname of your local server. Please refer to the Proteome Discoverer user manual for full instructions.
Bioworks
Note: Bioworks is no longer developed by Thermo. The instructions below are correct for Bioworks 3.3 SP1 but have not been tested with recent versions of Mascot Server.
Support for submitting searches direct to a Mascot Server was added to Thermo’s Bioworks in version 3.2, but we advise using Bioworks 3.3 SP1 to avoid some known issues with the first release. Mascot Server must be version 2.1 or later.
In Bioworks browser, choose Configuration off the Options menu. In the dialog, select Mascot Search and enter the Mascot Server URL in the form http://ec-vm2/mascot/cgi where ec-vm2 is replaced by the hostname of your local server.
When a data file is loaded, you can choose Mascot off the Actions menu to submit a search. Bioworks creates and saves an mzData format peak list for submission to Mascot.
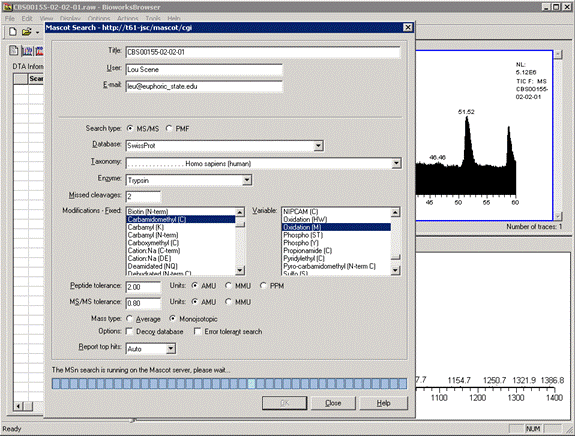
When the search is complete, you can load the Mascot results report in a web browser or download the results file to the Bioworks PC. Note that Bioworks has been superseded by Proteome Discoverer, and is no longer available.
Utilities
RawFileReader
RawFileReader is Thermo’s latest raw file access library.
MSFileReader
MSFileReader (download requires registration) is an older version of Thermo’s raw file access library. It is required by Mascot Distiller, and is installed automatically as part of a Mascot Distiller installation.
DeconMSn
DeconMSn has been developed at Pacific Northwest National Laboratory. It requires Xcalibur and Microsoft .NET 1.1 or later to be installed. It is not clear whether it can be made to run stand-alone, on a system without a full installation of Xcalibur.
DeconMSn can output either DTA or MGF peak lists. With high resolution data, parent monoisotopic mass is calculated using a modified THRASH approach. For low-resolution data, DeconMSn uses a support-vector machine based charge-detection algorithm to determine parent mass.
MSConvert
MSConvert is a component of ProteoWizard that converts between various file formats. It has a large number of options and can be configured as a data import filter in Mascot Daemon. Note that the input file format is not specified; msconvert auto-detects the format.
Acknowledgements
Sequest is a registered trademark of the University of Washington. Xcalibur is a registered trademark and Bioworks and Proteome Discoverer are trademarks of Thermo Electron Corporation.
