Mascot Distiller Toolboxes
Mascot Distiller functionality is organised in toolboxes.
Daemon Toolbox
The Mascot Daemon Toolbox enables the Mascot Distiller libraries to be called from Mascot Daemon, to process batches of data files and submit searches to a Mascot server. Peak lists are saved to disk automatically, and Distiller project files can also be created, allowing the raw data and peak list(s) to be opened in Distiller by clicking on a hyperlink in Daemon. With the Search Toolbox, search results will also be saved to be Distiller project file. With the Quantitation Toolbox, all stages of a quantitation experiment can be batch automated, form peak picking to saving the Distiller project complete with a quantitation report. (Note: Mascot Daemon is a component of the Mascot Server package)
Search Toolbox
The Search Toolbox is a collection of tools for protein identification and characterisation:
- Display Mascot Search results within Mascot Distiller
- Automatic de novo sequencing
- Automatic sequence tag calling
- Manual de novo and sequence tag
- Predict fragment ions from a peptide sequence and overlay them on an MS/MS spectrum
- Predict mass values from a protein digest and overlay them on a spectrum
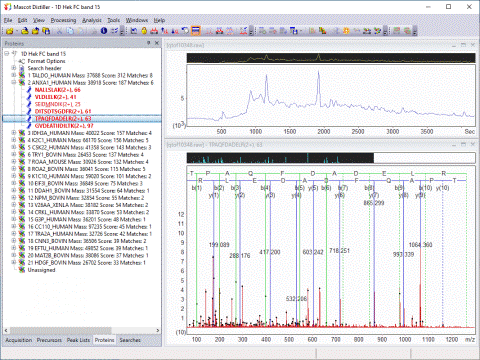
In the case of an MS/MS search, the top ranking peptide matches for each peak list are displayed under a Mascot Search Results node in DataSet Explorer. Each match is labelled with the peptide sequence and Mascot peptide score.
The Search Toolbox includes a powerful de novo sequencing algorithm:
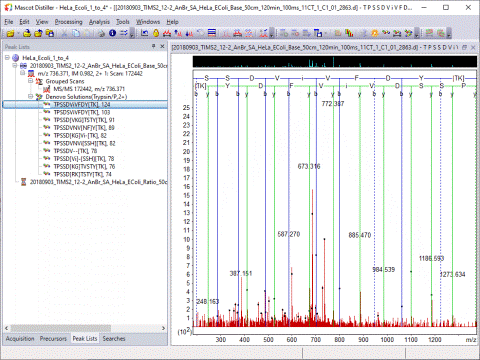
Quantitation Toolbox
Implements quantitation based on the relative intensities of extracted ion chromatograms for precursors in survey scans. This approach can be used “label-free”, or with any chemistry that creates a precursor mass shift, for example, 18O, AQUA, ICAT, ICPL, Metabolic labelling, and SILAC.
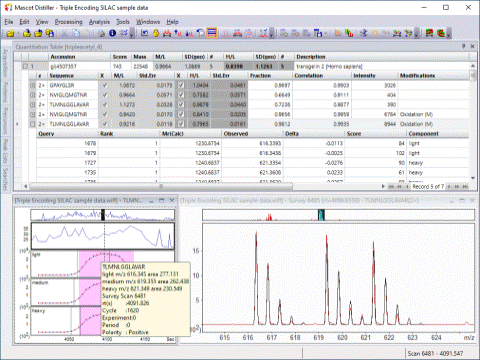
Individual ratios are automatically accepted or rejected by means of user defined quality thresholds. Corrections for isotope impurity and overlapping distributions are built in and overlays show how well the experimental peaks fit to the ideal distributions. The quantitation results can be exported as HTML, CSV and various graphical reports.
Developer Toolbox
The Developer Toolbox contains the documentation and support to develop and run programs that call the Mascot Distiller libraries. Distiller uses COM, so can be called from most Microsoft Windows programming languages. The Developer Toolbox provides a uniform Application Programmer Interface to all of the different raw file formats, greatly reducing development time.
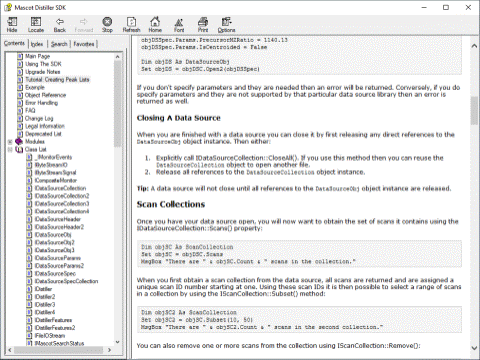
A run-time licence is required to execute the programs you develop on a PC which does not have Mascot Distiller Workstation installed.

