Identifying Mycobacteria with custom spectral libraries
Mass spectrometry can be used for rapid identification of difficult to culture or slow growing bacteria so that the patient can receive prompt treatment. The quickest identification method is searching the MS/MS spectra against a species-specific spectral library (SL). Here, we detail the steps to build a custom Mycobacteria spectral library and using it with Mascot Server.
Identification of Mycobacteria
One of the bacterial groups that falls into the difficult camp is Mycobacteria, a genus of ~195 rod shaped Gram-positive bacteria with thick, hydrophobic, and mycolic acid-rich cell walls made of peptidoglycan and arabinogalactan. Tuberculosis (M. tuberculosis) and leprosy (M. leprae) are human pathogens. Other Mycobacteria that do not cause tuberculosis or leprosy are known as nontuberculous mycobacteria or NTM.
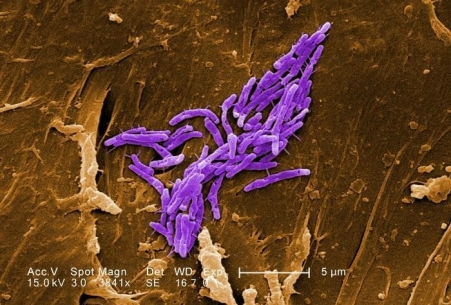
Mycobacterium fortuitum at 3841X magnification by SEM and digitally colorized.
The gold standard for mycobacterial detection starts with liquid culture in a mycobacterial growth indicator tube (MGIT), followed by sub-culturing on solid media. Species identification can be carried out by MALDI-TOF MS proteome or DNA fingerprinting. Due to the nature of Mycobacteria’s tough cell wall, genetic mutations to drug targets, and enzymes that modify or pump out drugs, they are often multi-drug resistant (MDR-TB) and extensively drug-resistant (XDR-TB). This complicates treatment as you need to identify the Mycobacteria’s strain and known drug resistances first.
Chinmaya Kotimoole et al. performed an analysis of proteomes of 7 NTM species totaling more than 20 million peptide spectrum matches gathered from 26 proteome data sets. The goal was to create a spectral library of peptide that are unique to individual species and strains to allow their rapid identification. A variety of instruments and search engines, including Mascot Server, were used to analyze or reanalyze DDA data. Final spectral libraries were created in Skyline in the BLIB format.
I selected a number of libraries from this meta analysis and converted them to MSP format using the blib2msp convertor introduced recently. Conversion time was a few minutes to 10 minutes depending on the size of the initial databases, which ranged from 20MB to 4GB.
| # | Full Species Name | MSP Library File |
|---|---|---|
| 1 | Mycobacteroides abscessus | M_abscessus_MSV000085363_20251216.msp |
| 2 | Mycobacteroides abscessus | M_abscessus_MSV000085363_FragPipe_Search_SpecLib.msp |
| 3 | Mycobacteroides abscessus | PXD022644_M_abscessus_LFQ_Proteomics_112022.msp |
| 4 | Mycobacterium avium subsp. paratuberculosis | M_avium_paratuberculosis_ProteomeDB_PD_Search_011023.msp |
| 5 | Mycobacterium kansasii | M_kansassi_Total_Proteomics_Database_Search_080122.msp |
| 6 | Mycobacteroides fortuitum | M_fortuitum_TP_FragPipe_ProteomeDB_Search_SpecLib_Final.msp |
| 7 | Mycobacterium intracellulare | M_intracellulare_TP_FragPipe_ProteomeDB_search_SpecLib.msp |
| 8 | Mycobacterium marinum | PXD003766_M_marinum_LFQ_proteomics_FragPipe_Search.msp |
| 9 | Mycolicibacterium vaccae | PASS00954_M_vaccae_Cytoplasma_CID_LFQ_Proteomics_SpecLib_112222.msp |
After conversion, I used mass spec data from a different Mycobacteria project so that we were not searching the same data files that were used to create the libraries. A recent study by Duran Bao et al. identified unique peptides for MRM studies from different Mycobacteria species and strains. As part of this study there were a number of DDA data sets from different NTM species that were made available in PRIDE project PXD059923. The samples were collected from sputum, biopsies of skin, or swabs from infected wounds etc and cultured in a BSA containing media. This is the selection I used:
| Sample name | Expected species |
| MAB1.raw | Mycobacteroides abscessus |
| MAB10.raw | Mycobacteroides abscessus |
| MAV1.raw | Mycobacterium avium complex |
| MAV5.raw | Mycobacterium avium complex |
| MFO1.raw | Mycolicibacterium fortuitum |
| MGO1.raw | Mycobacterium gordonae |
| MINT1.raw | Mycobacterium interjectum |
| MKA1.raw | Mycobacterium kansasii |
| MMU1.raw | Mycolicibacterium mucogenicum |
| MSI1.raw | Mycobacterium simiae |
| MTB1.raw | Mycobacterium tuberculosis |
| MTB25.raw | Mycobacterium tuberculosis complex |
| MTB8.raw | Mycobacterium tuberculosis complex |
Both these samples and the ones used to generate the initial spectral libraries contained a mixture of human proteins, BSA and associated bovine proteins from the culture media, as well as multiple Mycobacteria. Because of this, the libraries are not really suitable for species identification without further processing.
I went back to a technique previously mentioned in our archives and filtered the SL by peptides unique to the species of interest using Unipept. I extracted all the peptide sequences from the SL, processed them as an assay in Unipept, then drilled down to the appropriate Mycobacteria species, copied the unique representative peptides and filtered the library so that it only contained those spectra. The scripts for extracting the peptide sequences from a library and filtering an MSP library are included in the blib2msp convertor (GitHub repository). This is essentially the process Chinmaya Kotimoole et al. used when preparing their libraries except they created a custom tool. Filtering is simple:
# Extract from MSP file perl extract_peptides.pl -i my_library.msp # Output: my_library_peptides.txt # Filter MSP file (auto-generates output filename) perl filter_msp.pl -i library.msp -p peptide_list.txt # Output: library_filtered.msp
I processed some of the Bao et al. data files with Mascot Distiller and searched Mascot Server using a combination of different Mycobacteria spectral libraries, Unipept filtered SL, Unipept filtered SL plus the PRIDE_Contaminiants database and PRIDE_Human database, the SL combined with a Uniprot Mycobacteriaceae FASTA file (organism_id:1762), and just the Uniprot Mycobacteriaceae FASTA. There is no need to include modifications when using a library as they are included in the original analysis and library generation. For searches that included a FASTA file, I used fixed Carbamidomethyl (C) and variable Acetyl (Protein N-term), Oxidation (M). Peptide mass tolerance was ±20 ppm while Fragment mass tolerance was ±0.6 Da. Search times when run on a medium Mascot Server license, using 16 cores, varied depending on the size of the sample file and ranged from 1 to 5 minutes against an all spectral multi-library search and from 3 minutes to 11 minutes when including the Uniprot Mycobacteriaceae FASTA file.
Results
Searches against the combined SL were successful, although the actual results compared to expected species left a lot to be desired.
Database searching reports the best match in the database with a p-value that can be tested for statistical significance. The SL searches against unfiltered SL had a lot of matches, but very few were unique peptides. Searches against the filtered SL had less matches but they tended towards the largest SL. Samples containing a species without an associated SL acted as a negative control, as we were not expecting to find the target species. I would need to convert, filter and search all the available SL in order to make sure I had sufficient unique peptides representing all the species in my SL in order to have a good chance of identifying the species as per the original publication.
Searches that included a database search of a FASTA file were a lot more encouraging. I exported the identified peptides from the Mascot report and passed them through Unipept for a Lowest Common Ancestor (LCA) report.
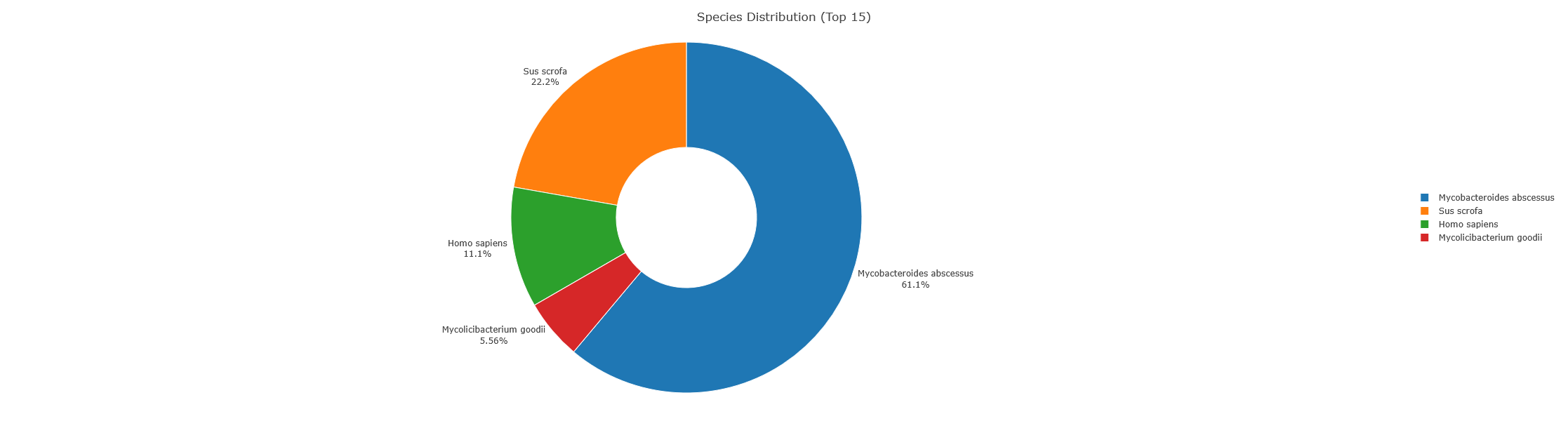
Sample MAB10.raw was easy to identify as Mycobacteroides abscessus. Of 218 total peptides with LCA assignments, 18 (8.3%) resolved to species level across 4 unique species. The remaining 200 (91.7%) peptides matched multiple organisms and were assigned to higher taxonomic levels (genus, family, order, etc.).
Top 5 Species – MAB10
- Mycobacteroides abscessus (species): 11 peptides (5.0%)
- Sus scrofa (species): 4 peptides (1.8%)
- Homo sapiens (species): 2 peptides (0.9%)
- Mycolicibacterium goodii (species): 1 peptides (0.5%)
For sample MSI1.raw there was no Mycobacterium simiae SL to search and there was only a single, presumably false positive, match to the largest SL Mycobacterium fortuitum. When the FASTA file was also searched, there were 330 peptides identified and Mycobacterium simiae plus its complex species group was the species the most unique peptides, 42 in all.
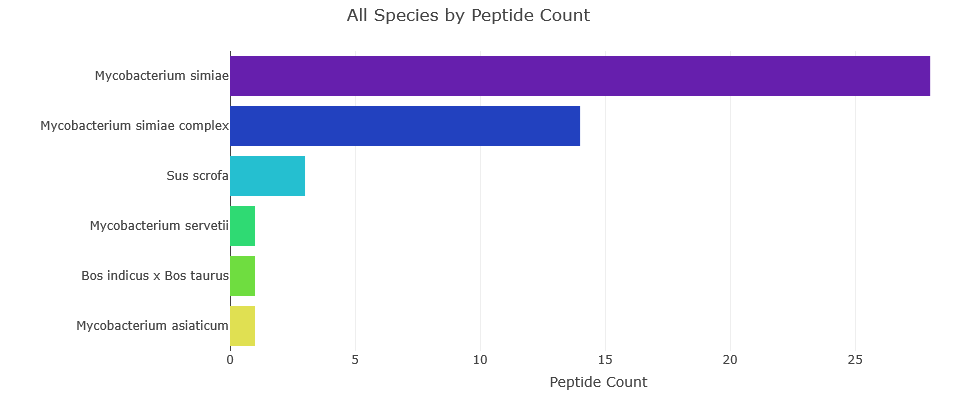
Top 5 Species – MSI1
- Mycobacterium simiae (species): 28 peptides (8.5%)
- Mycobacterium simiae complex (species group): 14 peptides (4.2%)
- Sus scrofa (species): 3 peptides (0.9%)
- Mycobacterium servetii (species): 1 peptides (0.3%)
- Bos indicus x Bos taurus (species): 1 peptides (0.3%)
The Mycobacterium avium complex was identified in sample MAV1.raw and MAB10.raw but no NTM species was identified from MAV5.raw. There were no unique Mycobacterium peptide matches to the MGO1.raw sample either. All the Mycobacterium tuberculosis samples were correctly identified as was Mycolicibacterium mucogenicum. Finally, the MKA1.raw sample had no unique peptide matches to Mycobacterium kansasii but did have unique peptide matches to Mycolicibacterium fluoranthenivorans.
The above is an example analysis with statistical evidence of the presence of the species in the sample. However, the absence of evidence does not prove that the expected species is not in the sample, just that no statistically significant evidence was found for it. The analysis is not equivalent to the PEP-TORCH algorithm but rather a simpler alternative.
Although the results from the spectral library searches themselves were not encouraging and would require a lot more libraries to be converted and filtered in order to make a representative library, the database and library searching worked as expected.
Keywords: metaproteomics, spectral library