New features in Mascot 2.0
Memory efficient mode for MudPIT-type searches
Very large searches are now divided into chunks, so that memory requirements are more modest. This is particularly important for 32 bit platforms, (Windows & Linux), which have an absolute limit of 2Gb per process, irrespective of the amount of RAM or swap space. This change also minimises any delay at the end of a large search performed on a Mascot Cluster.
Sequence tags, including error tolerant tags
Mascot now includes a powerful and complete implementation of the Mann & Wilm sequence tag. Ambiguous and error tolerant tags can be searched, and multiple tags can be specified for each query. Tags are scored probabilistically, and significance thresholds or expectation values used to avoid false positives. Syntax information is here.
Result reports now use Mascot Parser
Mascot result reports are generated on the fly by Perl scripts. To speed up report generation, and minimise memory requirements, most of the processing is now performed by the Mascot Parser library. This also makes the scripts simpler, so easier to customise.
Mascot Daemon Enhancements
Mascot Daemon now has the capability to call the Mascot Distiller library to process binary files in batch or real-time monitor mode, (requires additional licence).
Mascot Daemon has been split into two components: a user interface and a Windows service. This avoids the need for the application to be kept running on the desktop at all times. Communication between the service and a Mascot Server is fully asynchronous. There is no requirement to keep the connection between client and server open continuously.
Other changes to Mascot Daemon include comprehensive, context-sensitive help:
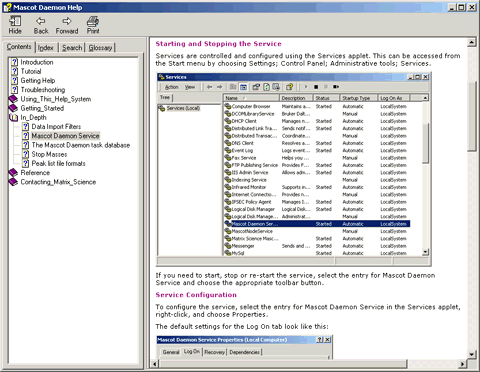
Support for d, v, and w ions
Side-chain cleavage ions d, v, and w can now be matched during MS/MS searches. These series are included in the default INSTRUMENT configuration for MALDI-TOF-TOF.
Other important changes
- Ambiguous amino acid residues X, B and Z are now explicitly iterated to find best match.
- Web pages updated and redesigned to eliminate frames.
- Nucleic acid translation now uses taxonomy to determine correct genetic code.
- Database maintenance utility has simplified interface.
- Windows 2003 Server, IIS 6, and Apache 2.x are fully supported by the installation program.
