New features in Mascot Server 2.8
New statistical model for error tolerant searches
The Error Tolerant Search finds unsuspected modifications, peptides with non-specific cleavage and residue substitutions. Mascot now assigns expect values to error tolerant (ET) matches based on statistical significance. If you submit the ET search as a target-decoy search, Mascot additionally estimates the false discovery rate (FDR). Proteins are selected for the ET pass based on at least one significant peptide match at the selected PSM FDR, by default 1% FDR. When the search has finished, the Protein Family Summary report chooses a significance threshold for ET matches that yields the target PSM FDR for the combined data set. Another improvement is that only queries that have no significant first pass match are selected for error tolerant searching.
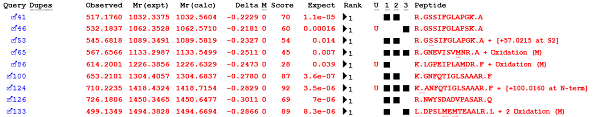
Increased Percolator sensitivity
The sensitivity of target-decoy database searches has been significantly improved when search results are re-ranked using Percolator. Mascot 2.8 ships with a new set of Percolator features, and Percolator has been updated to version 3.05. Depending on the search, the new feature set can give up to 100% improvement in sensitivity at 1% PSM FDR compared to Mascot 2.7. Retention time can be used as a feature if available. It is now possible to select a target FDR for PSMs or peptide sequences for Percolated results with the click of a button.
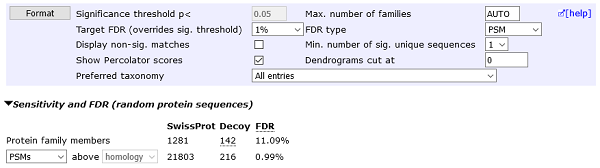
MS/MS searches are faster
MS/MS database searches are 20-35% faster than Mascot 2.7. Previously singly threaded file processing steps have been multithreaded. The improvement applies to searches of all sizes and hard disk types, whether HDD, SSD or RAID.
The speed of intralink and interlink searches with only a few protein accessions has been greatly improved. Additionally, memory usage has been greatly reduced when the search has more than a few dozen crosslinked proteins.
Other important changes
- Protein Family Summary report is now the default
- The Protein Family Summary report was introduced in Mascot 2.3, and it was the default report for large MS/MS searches. It is now the default report for all MS/MS searches. You can still view small MS/MS searches in Peptide Summary and Select Summary, but we recommend making best use of the rigorous protein inference in Protein Family Summary.
- Default false discovery rate for PSMs
- When you submit a target-decoy search, you can now select the target FDR for PSMs. At the end of the search, the Protein Family Summary report automatically selects a significance threshold that yields the closest FDR.
- Controls for rank selection for Percolator
- Two new mascot.dat options control which ranks are exported to Percolator. The default rule is: rank 1 plus further ranks from target database if score > 20 and difference from rank 1 score < 20%.
- Percolator is multithreaded
- Percolator 3.05 shipped with Mascot has been built with OpenMP enabled. The training step is now multithreaded. Additionally, creating the Percolator input (pip) file has been multithreaded.
- Configuration editor for crosslinking methods
- The Configuration Editor has been extended and now has a user interface for adding, copying, editing and viewing crosslinking methods.
- Export crosslinked matches in CSV and XML format
- Crosslinked search results can now be exported in CSV and XML format. The exported data contains monolink, looplink and crosslinked matches as well as linear peptide matches.
- Speed improvements to mzIdentML export
- Exporting search results as mzIdentML no longer exports site analysis as default, which makes exporting an order of magnitude faster. Site analysis can be enabled from the export form.
- Monitor handles test search timeout gracefully
- When Mascot Monitor brings a database online, it runs a test search. The test search can no longer time out, and the MonitorTestTimeout parameter has been removed.
And, of course, an embarrassing number of bug fixes.
