Processing Thunder-DDA-PASEF data with Mascot Distiller
Human leukocyte antigen (HLA) class I peptide ligands (HLAIps) are key targets in vaccine and immunotherapy development. They are also challenging to identify in standard mass-spec based proteomics; they are typically short, present at low abundance, and are not generated by a specific protease, requiring that search without enzyme specificity is conducted – dramatically increasing the search space compared with a standard Trypsin digest.
To address some of these challenges, Thunder-DDA-PASEF [1] was recently introduced as an optimized acquisition strategy for LC-IMS-MS on timsTOF instruments. The method modifies the standard DDA-PASEF workflow to better capture the characteristics of HLAlps, including the frequent presence of singly charged precursors and their narrow length distribution. The method takes its name from the lightning shape of the fragment isolation filter applied in the 1/K0 ion mobility and m/z channels.
DDA-PASEF files can be processed in Mascot Distiller, and Thunder-DDA-PASEF data are no exception. Using machine learning rescoring with MS2PIP and Percolator, we can recover many real HLA and no-enzyme matches which would otherwise fail to be significant due to the larger search space.
Example
To demonstrate this, we took several data files from the JPOST repository associated with the paper (PXD040385), half of which were captured using the standard DDA-PASEF method, and half the Thunder-DDA-PASEF method. Unfortunately, the archive does not contain the raw files used to generate figure 2 in the paper. However, it does contain files comparing the same plasma sample between the standard and thunder DDA-PASEF methods from another experiment which is not reported in the paper or supplementary data, three each for the Standard and Thunder DDA-PASEF settings. We reprocessed and searched those files using Mascot Distiller and Mascot Server 3.1.
We used the default processing options supplied for timsTOF files with Mascot Distiller with one change. The processing options include a 1/K0 tolerance on the precursor grouping defined by the
<precursorIonMobilityGroupTol>
element in the processing options file. By default, this is set to 0.026, but for these data a tighter tolerance of 0.01 was used. The setting is currently hidden in Distiller but it can be edited in a text editor – in the next release of Distiller, it will be available to edit in the GUI.
Search settings were derived from the paper, and the database searched was the mixed species database referred to in the paper. Searches were carried out using no enzyme specificity, and results were rescored using the standard Percolator features and also with the MS2PIP timsTOF2024 model shipped as part of the MS2Rescore package included with Mascot 3. The significance thresholds were adjusted for a 1% sequence FDR, and results are summarised in table 1 below:
| Significant sequences (1% FDR) | ||
|---|---|---|
| Isolation filter | Percolator | Percolator+timsTOF2024 |
| Standard | 3337 | 4937 |
| Thunder | 6378 | 8814 |
Table 1: Counts of significant peptide sequences at 1% FDR using the Standard and Thunder isolation filters, and re-processing with and without the timsTOF2024 model
So, we can see that the Thunder-DDA-PASEF files are giving us a large increase in significant sequences at 1% FDR compared to the standard method, and in both cases including predictions from the timsTOF2024 model is giving a >40% boost in significant sequences identified for both methods. All further comparisons were carried out using Percolator+timsTOF2024.
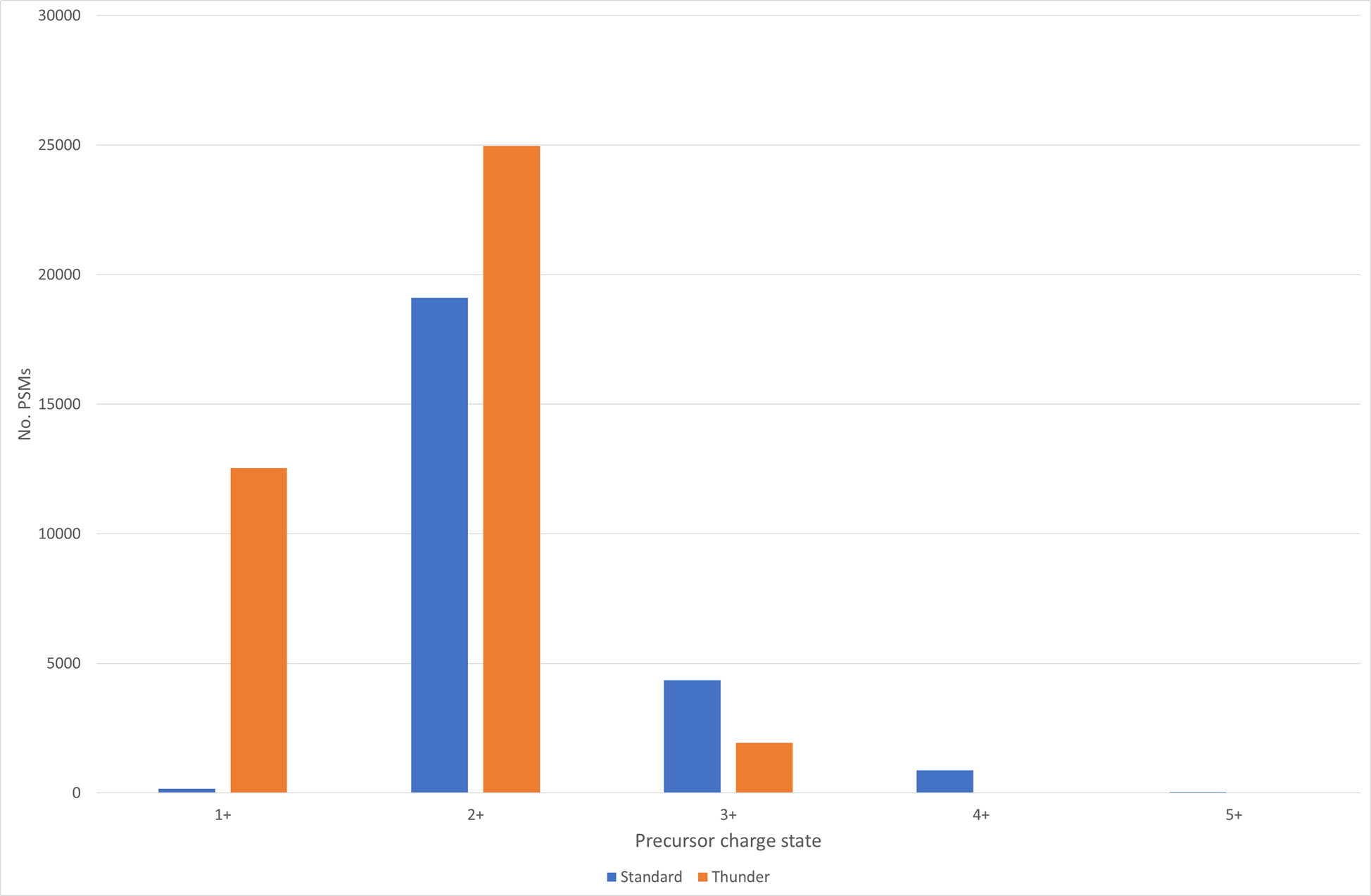
The Thunder DDA-PASEF isolation scheme was designed to improve the capture of peptide ions which match the expected profile of HLAIps, for example to include more singly charged peptides. Figure 1 below shows the comparison of the charge state distribution of significant PSMs between the standard and Thunder isolation schemes:
Figure 1: Charge state distribution for significant PSMs from Standard and Thunder DDA-PASEF methods at 1% sequence FDR
As you can see, the Thunder isolation scheme has not only restricted the charge state to 3+, but the number of significant matches to 1+ precursors has increased by ~82 fold. This can also be seen in the peptide length distribution – while the percentage of peptides identified with a length between 8-13 is similar for the two methods at roughly 90%, which is not unexpected given that 2+ is the most common charge state for both methods, the maximum length identified using Thunder isolation was 23 compared with 64 for the standard isolation pattern.
The authors estimated the number of HLAIps identified using NetMHCpan [2]. We took all the significant peptide sequences of lengths between 8 and 13 amino acid residues identified by the standard and Thunder isolation methods and ran them through NetMHCpan-4.1. Results are summarised in table 2 below:
| Isolation filter | No. predicted HLAIps |
|---|---|
| Standard | 2919 |
| Thunder | 4860 |
Table 2: Counts of peptides length 8-13 identified in the Standard and Thunder DDA-PASEF search results predicted to be HLAIps by NetMHCpan
As you can see, the results from the Thunder-DDA-PASEF isolation scheme show a significant increase in the number of predicted HLAIps.
Overall, the use of Mascot Distiller to process the data, Mascot Server 3.1 to search the generated peaklists, refine the results using machine learning and including spectral prediction information from MS2PIP are giving excellent results with these data.
References:
- Gomez-Zepeda D, Arnold-Schild D, Beyrle J, et al. Thunder-DDA-PASEF enables high-coverage immunopeptidomics and is boosted by MS2Rescore with MS2PIP timsTOF fragmentation prediction model. Nat Commun. 2024;15(1):2288. Published 2024 Mar 13. doi:10.1038/s41467-024-46380-y
- Hoof I, Peters B, Sidney J, et al. NetMHCpan, a method for MHC class I binding prediction beyond humans. Immunogenetics. 2009;61(1):1-13. doi:10.1007/s00251-008-0341-z
Keywords: hla, machine learning, Mascot Distiller, MS2PIP, peak picking, timsTOF