New features in Mascot 2.4
Database Manager
A new, browser-based Database Manager replaces the separate configuration and download scripts used in Mascot 2.3 and earlier. Up-to-date configuration information for popular public databases is maintained on the Matrix Science public web site. Provided the Mascot Server can connect to the Internet, these predefined database definitions will be kept up-to-date automatically.
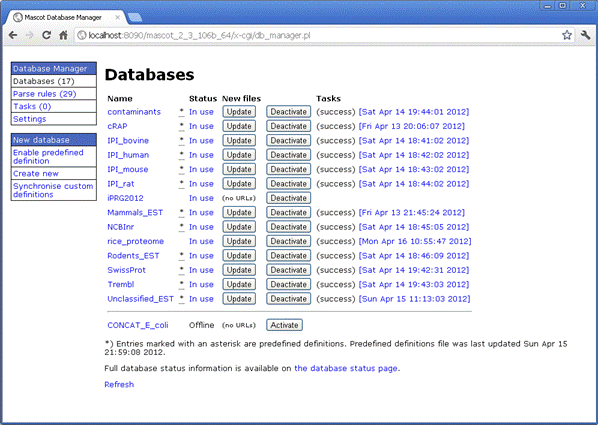
When you need to create custom definitions, configuration is more intuitive and more reliable. Updating a database or creating a schedule to check for updates every week or every month requires just a few mouse clicks
Report Builder
The second tab of the Protein Family Summary has been greatly enhanced and the name changed from Quantitation to Report Builder. Its main purpose is to build a customised table of protein hits, which is particularly useful if you need a minimal list of proteins for a publication. You can choose which columns to include and their order, filter out proteins that are of no interest, such as one-hit wonders, and export the table in CSV format directly to Excel.
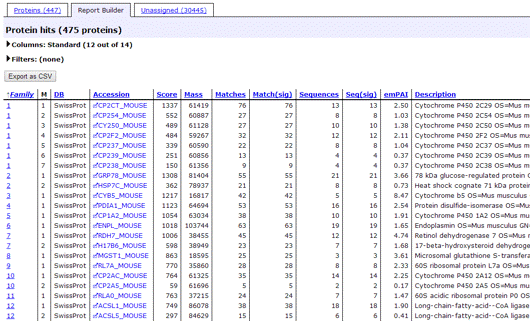
If the search included quantitation using Reporter or Multiplex protocols, protein level quantitation information is available. The filters make it easy to create a table in which, for example, proteins are listed if they are significantly up-regulated in one channel and significantly down-regulated in another.
Preferred taxonomy
If you are mainly interested in one particular taxonomy, but search a broader taxonomy to get hits to homologous or contaminant proteins, the preferred taxonomy control at the top of the Protein Family Summary can be used to promote the taxonomy of interest. For example, you might search green plants but want to see matches to maize where a family member contains a choice of same set proteins.
Target/Decoy search improvements
In Mascot 2.3 and earlier, you had to use trial and error to find the significance threshold that gave the required FDR. Slightly tedious. In Mascot 2.4, you can click a button at the top of the Protein Family Summary and have it go to the required FDR automatically
There is now a choice of algorithms for creating the decoy sequences in an automatic target/decoy search:
- Randomised proteins (as in 2.3)
- Reversed proteins
- Randomised peptides
Site analysis for variable modifications
Bernhard Kuster has shown how the Mascot score difference can be used for reliable site assignment when there are matches to the same spectrum with different arrangements of modifications. (Confident Phosphorylation Site Localization Using the Mascot Delta Score.) We have implemented this approach in the Mascot 2.4 Peptide View report. The only major difference is that site analysis works for any variable modification, not just phosphorylation. Site assignment will be displayed automatically whenever the top match is significant and multiple arrangements are possible for one or more variable mods.
| Score | Mr(calc) | Delta | Sequence | Site Analysis |
|---|---|---|---|---|
| 83.4 | 1846.7179 | 0.1889 | DIGSESTEDQAMEDIK | Phospho S4 84.56% |
| 75.8 | 1846.7179 | 0.1889 | DIGSESTEDQAMEDIK | Phospho S6 14.73% |
| 62.7 | 1846.7179 | 0.1889 | DIGSESTEDQAMEDIK | Phospho T7 0.72% |
| 26.9 | 1846.7808 | 0.1261 | KLNSNPENYCESELK | |
| 22.8 | 1846.7729 | 0.1339 | KMEDSVGCLETAEEVK | |
| 15.5 | 1846.9230 | -0.0161 | GAYTIEQHPVLGLEIK | |
| 14.2 | 1846.7729 | 0.1339 | KMEDSVGCLETAEEVK | |
| 13.9 | 1846.8754 | 0.0315 | YVKGIYENLPSIDEK | |
| 13.8 | 1846.8866 | 0.0202 | QLIEAPDPVPSFEVAR | |
| 13.3 | 1846.9052 | 0.0016 | KIDFSNIAMLFGGVQK |
Other important changes
- ActiveState Perl versions 5.8 to 5.14 (inclusive) are supported
- Additional parameters added to MGF format, such as RAWFILE and LOCUS
- Re-centroiding of peak list restricted to cases where data is clearly profile
- Comment lines allowed in Fasta files
- Text and number search within Protein Family Summary enhanced
- Satellite ions no longer restricted to fragments containing R
